
| Sequence ID | dm3.chrX |
|---|---|
| Location | 8,057,778 – 8,057,873 |
| Length | 95 |
| Max. P | 0.636167 |

| Location | 8,057,778 – 8,057,873 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 66.15 |
| Shannon entropy | 0.65953 |
| G+C content | 0.57130 |
| Mean single sequence MFE | -29.97 |
| Consensus MFE | -13.40 |
| Energy contribution | -14.83 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.38 |
| Mean z-score | -0.90 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.636167 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 8057778 95 + 22422827 CGGGCCAACGGUAAAGCACAACAAAGUGUGGUCACCACUACGAAUUGGGCCAGAAAUCGGUGGCCAAGUGGAUCUGGCAUGGAGAUGGCGACGAA ((.((((.(......((((......)))).(((((((((........(((((........))))).)))))...)))).....).))))..)).. ( -26.60, z-score = 0.15, R) >droEre2.scaffold_4690 10429285 93 + 18748788 -CGGCCAACGGUAAAGCAGCACCAACUGCG-CCACCACUUCGAAUUGGGCCAGAAAUCGGUGGCCAAGUGCAGCUGGAGUGGAGGUGGAGGCGAG -..(((..(......((((......))))(-((.((((((((.(((.(((((........))))).))).....)))))))).))))..)))... ( -34.10, z-score = -0.74, R) >droYak2.chrX 8518261 74 - 21770863 CUGGCCAACGGUAAAG----------UGUGG-CACCACUACGAAUUGGGCCAGAAAUCGGUGACCAAGUGGCGCUGGCUACGAAA---------- ((((((..((((...(----------((((.-...))).))..))))))))))...((((((.((....))))))))........---------- ( -24.30, z-score = -0.96, R) >droSec1.super_44 143956 95 + 251150 CAGGCCAACGGUAAAGCACAACAAAGUGCGGCCACCACUACGAAUUGGGCCAGAAAUCGGUGGCCAAGUGGAUCUGGCAUGAAGGUGGCGACGCA ((.((((..(((...((((......)))).))).(((((........(((((........))))).)))))...)))).))...(((....))). ( -34.00, z-score = -1.60, R) >droSim1.chrX_random 2415371 95 + 5698898 CAGGCCAACGGUAAAGCACAACAAAGUGCGGCCACCACUGCGAAUUGGGCCAGAAAUCGCUGGCCAAGUGGAUCUGGCAUGGAGGUGGCGACGCA ((.((((..(((...((((......)))).))).(((((........((((((......)))))).)))))...)))).))...(((....))). ( -38.70, z-score = -2.16, R) >dp4.chrXR_group6 6066568 72 + 13314419 ----------GUAAAGUGCG-------GCAGCUCGUGCUCCAAAUGGUGCUCCAAAUGAUGAUCCAAGUGGGGCAGGAUACCGGCCUGU------ ----------.........(-------((...((.(((((((..(((..(((.....)).)..)))..))))))).)).....)))...------ ( -22.10, z-score = -0.09, R) >consensus CAGGCCAACGGUAAAGCACAACAAAGUGCGGCCACCACUACGAAUUGGGCCAGAAAUCGGUGGCCAAGUGGAGCUGGCAUGGAGGUGGCGACG_A ...((((..(((...((((......)))).))).(((((........(((((........))))).)))))...))))................. (-13.40 = -14.83 + 1.44)



| Location | 8,057,778 – 8,057,873 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 66.15 |
| Shannon entropy | 0.65953 |
| G+C content | 0.57130 |
| Mean single sequence MFE | -25.64 |
| Consensus MFE | -11.25 |
| Energy contribution | -11.28 |
| Covariance contribution | 0.03 |
| Combinations/Pair | 1.46 |
| Mean z-score | -0.84 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.529077 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 8057778 95 - 22422827 UUCGUCGCCAUCUCCAUGCCAGAUCCACUUGGCCACCGAUUUCUGGCCCAAUUCGUAGUGGUGACCACACUUUGUUGUGCUUUACCGUUGGCCCG ......((((.(.....((((((.....((((...))))..))))))............(((((.((((......))))..)))))).))))... ( -23.50, z-score = -0.65, R) >droEre2.scaffold_4690 10429285 93 - 18748788 CUCGCCUCCACCUCCACUCCAGCUGCACUUGGCCACCGAUUUCUGGCCCAAUUCGAAGUGGUGG-CGCAGUUGGUGCUGCUUUACCGUUGGCCG- ...(((........(((.((((((((.....((((((.(((((.((......))))))))))))-))))))))).).))..........)))..- ( -30.07, z-score = -0.70, R) >droYak2.chrX 8518261 74 + 21770863 ----------UUUCGUAGCCAGCGCCACUUGGUCACCGAUUUCUGGCCCAAUUCGUAGUGGUG-CCACA----------CUUUACCGUUGGCCAG ----------.......(((((((((....)))..........(((((((........))).)-)))..----------.......))))))... ( -20.70, z-score = -1.14, R) >droSec1.super_44 143956 95 - 251150 UGCGUCGCCACCUUCAUGCCAGAUCCACUUGGCCACCGAUUUCUGGCCCAAUUCGUAGUGGUGGCCGCACUUUGUUGUGCUUUACCGUUGGCCUG ......((((.(.....((((...(((((.(((((........)))))........))))))))).((((......))))......).))))... ( -28.40, z-score = -0.81, R) >droSim1.chrX_random 2415371 95 - 5698898 UGCGUCGCCACCUCCAUGCCAGAUCCACUUGGCCAGCGAUUUCUGGCCCAAUUCGCAGUGGUGGCCGCACUUUGUUGUGCUUUACCGUUGGCCUG ...(((((((((....(((..((.......((((((......))))))....)))))..)))))).((((......)))).........)))... ( -31.80, z-score = -1.07, R) >dp4.chrXR_group6 6066568 72 - 13314419 ------ACAGGCCGGUAUCCUGCCCCACUUGGAUCAUCAUUUGGAGCACCAUUUGGAGCACGAGCUGC-------CGCACUUUAC---------- ------...(((.(((.((.(((.(((..(((.((........))...)))..))).))).)))))))-------).........---------- ( -19.40, z-score = -0.69, R) >consensus U_CGUCGCCACCUCCAUGCCAGAUCCACUUGGCCACCGAUUUCUGGCCCAAUUCGUAGUGGUGGCCGCACUUUGUUGUGCUUUACCGUUGGCCUG .................(((((........(((((........))))).........(((.....)))...................)))))... (-11.25 = -11.28 + 0.03)


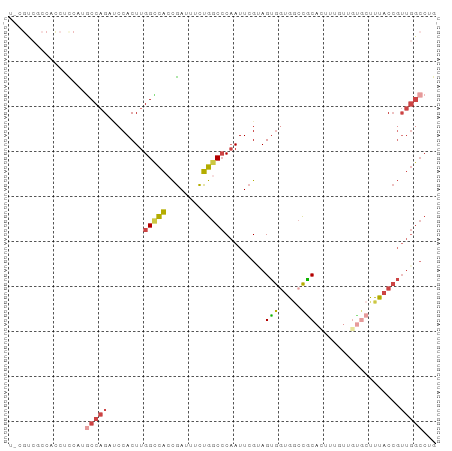
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:25:04 2011