
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 10,572,297 – 10,572,390 |
| Length | 93 |
| Max. P | 0.621744 |

| Location | 10,572,297 – 10,572,390 |
|---|---|
| Length | 93 |
| Sequences | 10 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 66.60 |
| Shannon entropy | 0.66880 |
| G+C content | 0.47190 |
| Mean single sequence MFE | -26.15 |
| Consensus MFE | -10.24 |
| Energy contribution | -9.84 |
| Covariance contribution | -0.40 |
| Combinations/Pair | 1.37 |
| Mean z-score | -1.17 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.621744 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 10572297 93 - 23011544 UGAUGGUGGAGUCGAAGAGUUGGAGA--------GUCGAGGCCCAAGGCUCGAACCCUUAAUUAUAUUGAGUGUAUGAAAUCC-UUUGGCCACACAAUGGCC------- .....(((..(((((((..((.(((.--------((((((.(....).)))).)).)))..((((((.....))))))))..)-))))))..))).......------- ( -24.60, z-score = -0.63, R) >droSim1.chr2L 10376177 101 - 22036055 UGAUGGUGGAGUCGUAGAGUUGGAGAGUUUGGGAGUCGAGGCCCAAGGCUCGAACCCUUAAUUAUAUUGAGUGUAUGAAAUCC-UUUGGCCACACAAUGGCC------- .......(((.(((((.(.(..(..((((.(((..(((((.(....).)))))..))).))))...)..).).)))))..)))-...(((((.....)))))------- ( -30.70, z-score = -1.43, R) >droSec1.super_3 5998425 101 - 7220098 UGAUGGUGGAGUCGUAGAGUUGGAGAGUUUGAGAGUCGAGGCCCAAGGCUCGAACCCUUAAUUAUAUUGAGUGUAUGAAAUCC-UUUGGCCACACAAUGGCC------- .......(((.(((((.(.(..(.....(((((.((((((.(....).)))).)).))))).....)..).).)))))..)))-...(((((.....)))))------- ( -27.50, z-score = -1.03, R) >droYak2.chr2L 6981715 84 - 22324452 -------GAAGU--AGAAGUUGGAGA--------GUCGAGGCCCAAGGCUCGAACCCUUAAUUAUAUUGAGUGUAUGAAAUCC-UUUGGCCACACAAUGGCC------- -------.....--.......(((..--------.(((((.(....).)))))((.((((((...)))))).))......)))-...(((((.....)))))------- ( -19.20, z-score = -0.47, R) >droEre2.scaffold_4929 11786230 84 + 26641161 -------GGAG--AUGGAGUCGGAGA--------GUCGAGGCCGAAGGCUCGAAUCCUUAAUUAUAUUGAGUGUAUGAAAUCC-UUUGGCCACACAAUGGCC------- -------....--..(((...(((..--------.(((((.(....).))))).)))....((((((.....))))))..)))-...(((((.....)))))------- ( -23.50, z-score = -0.89, R) >droAna3.scaffold_12916 14422159 96 - 16180835 ----ACUCCGGGCGGACGAUUGGGCGGAUGGGUGGACG-GCCACAAGGCUUGAACCCUUAAUUAUAUUGAGUGUAUGAAAUCC-UUUGGCCUCACAAUGGCC------- ----..(((....))).........((((((((....(-(((....))))...))))....((((((.....)))))).))))-...((((.......))))------- ( -26.10, z-score = 0.75, R) >droWil1.scaffold_180708 1786610 90 + 12563649 ------UGAGGAAGAAGGGGCAGGAUGAACAGGCUCAGGCUAUCAGGCCUUGAGCCCUUAAUUAUAUUGAGUGUAUGAAAUCCUUUUGGCCACACA------------- ------...((..(((((((...........((((((((((....)))).)))))).....((((((.....))))))..)))))))..)).....------------- ( -29.10, z-score = -1.62, R) >droVir3.scaffold_12963 9771538 92 + 20206255 ----------------CGUUUUGUGUGGCGCAUAACUUGGGCUCAAGGCUCAAGCCCUUAAUUAUAUUGAGUGUACGAAAUCC-UUUAGCCACACAAUGCAGCCCAGAC ----------------.((.(((((((((......(((((((.....)))))))..((((((...))))))............-....))))))))).))......... ( -28.70, z-score = -2.42, R) >droMoj3.scaffold_6500 13218776 92 + 32352404 ----------------UUUUUUGUGUGGCGCAUAACUUGGGCUCAAGGCUCAAGCCCUUAAUUAUAUUGAGUGUAUGAAAUCC-UUUAGCCACACAAUGCAGCCCAUCC ----------------....(((((((((......(((((((.....))))))).......((((((.....)))))).....-....)))))))))............ ( -27.00, z-score = -2.19, R) >droGri2.scaffold_15252 8699405 90 + 17193109 ---------------GAGUUUUGUAUGGCGUAUAACUUGGGCUCAAGGCUCAAGCCCUUAAUUAUAUUGAGUGUAUGAAAUCC-UUUAGCCACACAAUGCGCUCCA--- ---------------((((.((((.((((......(((((((.....))))))).......((((((.....)))))).....-....)))).))))...))))..--- ( -25.10, z-score = -1.80, R) >consensus _______GGAG__G_AGAGUUGGAGAG__G_AUAGUCGAGGCCCAAGGCUCGAACCCUUAAUUAUAUUGAGUGUAUGAAAUCC_UUUGGCCACACAAUGGCC_______ .......................................((((.((((.(((.((.((((((...)))))).)).)))...)).)).)))).................. (-10.24 = -9.84 + -0.40)


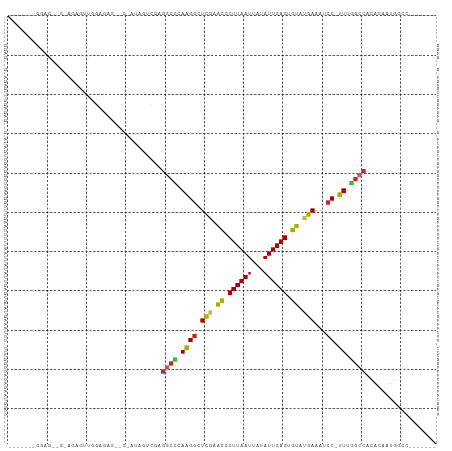
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:30:31 2011