
| Sequence ID | dm3.chrX |
|---|---|
| Location | 7,902,170 – 7,902,264 |
| Length | 94 |
| Max. P | 0.553421 |

| Location | 7,902,170 – 7,902,264 |
|---|---|
| Length | 94 |
| Sequences | 9 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 76.53 |
| Shannon entropy | 0.42219 |
| G+C content | 0.42019 |
| Mean single sequence MFE | -23.15 |
| Consensus MFE | -13.04 |
| Energy contribution | -13.22 |
| Covariance contribution | 0.18 |
| Combinations/Pair | 1.40 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.553421 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
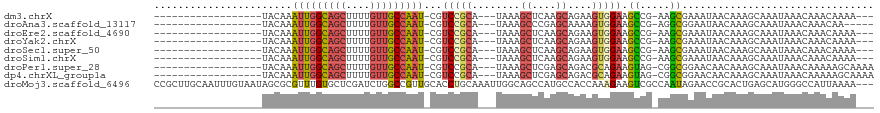
>dm3.chrX 7902170 94 - 22422827 ------------------UACAAAUUGGCAGCUUUUGUUGCCAAU-CGUCCGCA---UAAAGCUCAAGCAGAAGUGGAAGCCG-AAGCGAAAUAACAAAGCAAAUAAACAAACAAAA--- ------------------......((((((((....))))))))(-(((((((.---....((....))....)))))...))-).((...........))................--- ( -20.50, z-score = -1.60, R) >droAna3.scaffold_13117 2117872 92 - 5790199 ------------------UACAAAUUGGCAGCUUUUGUUGCCAAU-CGUCCGCA---UAAAGCCCGAGCAAAAGUGGAAGCCG-AGGCGGAAUAACAAAGCAAAUAAACAAACAA----- ------------------.....(((((((((....)))))))))-..(((((.---....(((((........)))..))..-..)))))........................----- ( -22.50, z-score = -1.41, R) >droEre2.scaffold_4690 16816151 94 - 18748788 ------------------UACAAAUUGGCAGCUUUUGUUGCCAAU-CGUCCGCA---UAAAGCUCAAGCAGAAGUGGAAGCCG-AAGCGAAAUAACAAAGCAAAUAAACAAACAAAA--- ------------------......((((((((....))))))))(-(((((((.---....((....))....)))))...))-).((...........))................--- ( -20.50, z-score = -1.60, R) >droYak2.chrX 8361560 94 + 21770863 ------------------UACAAAUUGGCAGCUUUUGUUGCCAAU-CGUCCGCA---UAAAGCUCAAGCAGAAGUGGAAGCCG-AAGCGAAAUAACAAAGCAAAUAAACAAACAAAA--- ------------------......((((((((....))))))))(-(((((((.---....((....))....)))))...))-).((...........))................--- ( -20.50, z-score = -1.60, R) >droSec1.super_50 186828 94 - 201529 ------------------UACAAAUUGGCAGCUUUUGUUGCCAAU-CGUCCGCA---UAAAGCUCAAGCAGAAGUGGAAGCCG-AAGCGAAAUAACAAAGCAAAUAAACAAACAAAA--- ------------------......((((((((....))))))))(-(((((((.---....((....))....)))))...))-).((...........))................--- ( -20.50, z-score = -1.60, R) >droSim1.chrX 6316955 94 - 17042790 ------------------UACAAAUUGGCAGCUUUUGUUGCCAAU-CGUCCGCA---UAAAGCUCAAGCAGAAGUGGAAGCCG-AAGCGAAAUAACAAAGCAAAUAAACAAACAAAA--- ------------------......((((((((....))))))))(-(((((((.---....((....))....)))))...))-).((...........))................--- ( -20.50, z-score = -1.60, R) >droPer1.super_28 305707 97 + 1111753 ------------------UACAAAUUGGCAGCUUUUGUUGCCAAU-CGUCCGCA---UAAAGCUCGAGCAGACGCAGAAGUAG-CGGCGGAACAACAAAGCAAAUAAACAAAAAGCAAAA ------------------.....(((((((((....)))))))))-..(((((.---....((....))...(((.......)-)))))))........((.............)).... ( -26.02, z-score = -2.34, R) >dp4.chrXL_group1a 6781815 97 + 9151740 ------------------UACAAAUUGGCAGCUUUUGUUGCCAAU-CGUCCGCA---UAAAGCUCGAGCAGACGCAGAAGUAG-CGGCGGAACAACAAAGCAAAUAAACAAAAAGCAAAA ------------------.....(((((((((....)))))))))-..(((((.---....((....))...(((.......)-)))))))........((.............)).... ( -26.02, z-score = -2.34, R) >droMoj3.scaffold_6496 4914423 117 - 26866924 CCGCUUGCAAUUUGUAAUAGCGCGUUUGUGCUCGAUCUGGCCGUUGCACCUGCAAAUUGGCAGCCAUGCCACCAAAGAAGUCGCCAAUAGAACCGCACUGAGCAUGGGCCAUUAAAA--- .(((((((.....)))..)))).(((..((((((...((((..((((....))))..(((((....)))))...........))))...(.....)..))))))..)))........--- ( -31.30, z-score = 0.89, R) >consensus __________________UACAAAUUGGCAGCUUUUGUUGCCAAU_CGUCCGCA___UAAAGCUCAAGCAGAAGUGGAAGCCG_AAGCGAAAUAACAAAGCAAAUAAACAAACAAAA___ .......................(((((((((....)))))))))......((........((....))..........((.....))...........))................... (-13.04 = -13.22 + 0.18)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:24:49 2011