| Sequence ID | dm3.chrX |
|---|---|
| Location | 7,738,762 – 7,738,842 |
| Length | 80 |
| Max. P | 0.823307 |

| Location | 7,738,762 – 7,738,842 |
|---|---|
| Length | 80 |
| Sequences | 12 |
| Columns | 83 |
| Reading direction | reverse |
| Mean pairwise identity | 85.87 |
| Shannon entropy | 0.30141 |
| G+C content | 0.44484 |
| Mean single sequence MFE | -23.94 |
| Consensus MFE | -17.67 |
| Energy contribution | -17.88 |
| Covariance contribution | 0.22 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.81 |
| SVM RNA-class probability | 0.823307 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
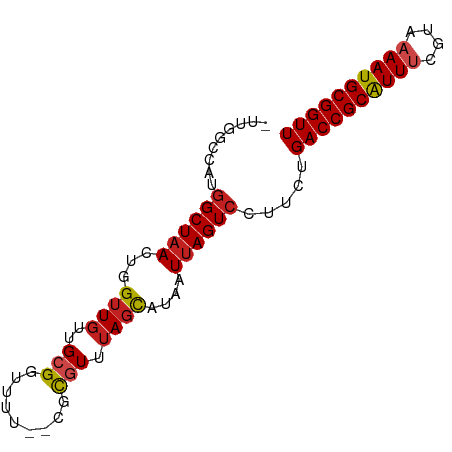
>dm3.chrX 7738762 80 - 22422827 -UUGGCCAUGGCUAACUGGUUGUUGCGGCUUU--CGCGUUUAGCAUAAUUAGUCUUUCUGACCGCAUUACGUAAAAUGCGGUU -.((((....))))(((((((((.((.((...--.)).....)))))))))))......(((((((((......))))))))) ( -24.20, z-score = -1.60, R) >droSim1.chrX 6191550 80 - 17042790 -UUGGCCAUGGCUAACUGGUUGUUGCGGCUUU--CGCGUUUAGCAUAAUUAGUCUUUCUGACCGCAUUUCGUAAAAUGCGGUU -.((((....))))(((((((((.((.((...--.)).....)))))))))))......((((((((((....)))))))))) ( -24.60, z-score = -1.91, R) >droSec1.super_50 27870 80 - 201529 -UUGGCCAUGGCUAACUGGUUGUUGCGGCUUU--CGCGUUUAGCAUAAUUAGUCUUUCUGACCGCAUUUCGUAAAAUGCGGUU -.((((....))))(((((((((.((.((...--.)).....)))))))))))......((((((((((....)))))))))) ( -24.60, z-score = -1.91, R) >droEre2.scaffold_4690 16653237 79 - 18748788 -UUGGCCAUGGCUAACUGGUUGUUGCGGUUUU--CGCGUUUAGCAUAAUUAGUCUUUC-GACCGCAUUUCGUAAAAUGCGGUU -.((((....))))(((((((((((((.....--))))......))))))))).....-((((((((((....)))))))))) ( -24.10, z-score = -1.94, R) >droAna3.scaffold_13117 4180547 78 + 5790199 ---GAACAUGGCUAACUGGUUGUUGCGGUUUU--GGCAUUUAGCAUAAUUAGUCCUUCUGACCGCAUUUCGUAAAAUGCGGUU ---(((...((((((...((((.(((......--.)))..))))....)))))).))).((((((((((....)))))))))) ( -21.60, z-score = -1.64, R) >droPer1.super_14 1244788 81 - 2168203 CUCGGCCAUGGCUAACUGGAUGUUGCGGUUAC--AACGUUUAGCAUAAUUAGUCCUUCUGACCGCAUUUCGUAAAAUGCGGUU .((((....((((((((((((((((......)--))))))))).....))))))...))))((((((((....)))))))).. ( -25.50, z-score = -2.49, R) >droYak2.chrX 8189592 79 + 21770863 -UUGACCAUGGCUAACUGGUUGUUGCGGUUUU--CGCGUUUAGCAUAAUUAGUCCUUU-GACCGCAUUUCGUAAAAUGCGGUU -........((((((...((((.((((.....--))))..))))....))))))....-((((((((((....)))))))))) ( -22.90, z-score = -1.83, R) >dp4.chrXL_group1e 164991 81 + 12523060 CUCGGCCAUGGCUAACUGGUUGUUGCGGUUAC--AACGUUUAGCAUAAUUAGUCCUUCUGACCGCAUUUCGUAAAAUGCGGUU ...(((....))).(((((((((.((......--........)))))))))))......((((((((((....)))))))))) ( -22.64, z-score = -1.48, R) >droWil1.scaffold_181096 2685751 80 + 12416693 -AUGGAAAUGGCUCACUGGUUGCUGCGGUUUU--GGCAUUUAGCAUAAUUAGUCCUUUCGACCGCAUUUCGUAUAAUGCGGUU -...((((.((...((((((((.(((.((...--.)).....)))))))))))))))))(((((((((......))))))))) ( -23.60, z-score = -1.73, R) >droVir3.scaffold_13042 2533518 82 - 5191987 -UCGGCCAUGGCUAAGUGGUUGUUGCGGUUUUACGCUGUUUAGCAUAAUUAGUCCAUUUGACCGCGUUUCGCGAAAUGCGGUU -......((((((((...((((..(((......)))....))))....)))).))))..((((((((((....)))))))))) ( -26.20, z-score = -1.35, R) >droMoj3.scaffold_6328 3398818 80 + 4453435 ---GGCCAUGGCUAAGUGGUUGUUGCGGUUUUACGCUGUUUAGCAUAAUUAGUCCUUUUGACCGCGUUUUGCGAAAUGCGGUU ---......((((((...((((..(((......)))....))))....)))))).....((((((((((....)))))))))) ( -24.30, z-score = -1.23, R) >droGri2.scaffold_15203 5373647 82 - 11997470 -UCCGUGCUGGCUAAGUGGUUUUUGCGGUUUUACACUGUUUAGUAUAAUUAGUACUUUUGACCGCGUUUUGUGAAAUGCGGUU -...((((((((((((..((..............))..)))))).....))))))....((((((((((....)))))))))) ( -23.04, z-score = -2.05, R) >consensus _UUGGCCAUGGCUAACUGGUUGUUGCGGUUUU__CGCGUUUAGCAUAAUUAGUCCUUCUGACCGCAUUUCGUAAAAUGCGGUU .........((((((...((((..(((.........))).))))....)))))).....((((((((((....)))))))))) (-17.67 = -17.88 + 0.22)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:24:29 2011