
| Sequence ID | dm3.chrX |
|---|---|
| Location | 7,489,543 – 7,489,631 |
| Length | 88 |
| Max. P | 0.734019 |

| Location | 7,489,543 – 7,489,631 |
|---|---|
| Length | 88 |
| Sequences | 7 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 74.03 |
| Shannon entropy | 0.47612 |
| G+C content | 0.39705 |
| Mean single sequence MFE | -19.67 |
| Consensus MFE | -11.44 |
| Energy contribution | -11.52 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.32 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.734019 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 7489543 88 - 22422827 CAUAAAGGUGAA---AAGGUAGCCGAGCGAUCAUCAAUGCCACUUGAUGGUCACACAACGAGAUAAAAA--AUGUGUGAAAU-UUCAGGUUCCU------- ............---.(((.((((....(((((((((......)))))))))(((((............--.))))).....-....)))))))------- ( -20.92, z-score = -1.60, R) >droSim1.chrX 6010148 91 - 17042790 CAUAAAGGUGAAAUAAAGGUAGCCGAGCGAUCAUCAAUGCCACUUGAUGGUCACACAACGAGAUAAAAA--AUGUGUGAAAU-UUCAGGUUUCA------- ........((((.....((((...((.......))..))))((((((...(((((((............--.)))))))...-.))))))))))------- ( -19.02, z-score = -1.00, R) >droSec1.super_24 696936 91 - 912625 CAUAAAGGUGAAAUAAAGGUAGCCGAGCGAUCAUCAAUGCCACUUGAUGGUCACACAACGAGAUAAAAA--AUGUGUGAAAU-UUCAGGUUCCA------- .................((.((((....(((((((((......)))))))))(((((............--.))))).....-....)))))).------- ( -20.92, z-score = -1.60, R) >droYak2.chrX 7932957 68 + 21770863 ----------------------CAUAAAGGUGA-AAAUGCCACUUGAUGGUCACACAACGAGAUAAAAA--AUGUGUGAAAU-UUCAGGUUCGC------- ----------------------......(((..-....)))((((((...(((((((............--.)))))))...-.))))))....------- ( -16.62, z-score = -2.28, R) >droEre2.scaffold_4690 16404410 88 - 18748788 CAUAAAGGUGGA---AAAGUAGCCGAGCGAUCAUCAAUGCCACUUGAUGGUCACACAACGAGAUGAAAA--AUGUGUGAAAU-UUCAGGUUCCU------- ......(((...---......)))(((((((((((((......)))))))))(((((............--.))))).....-.....))))..------- ( -19.22, z-score = -0.60, R) >dp4.chrXL_group1e 11488066 94 - 12523060 AACAAAGGCCAA---AAGGUCGCUGGGA----AUCAAUGCCACUUGAUGGUUACACAAAGUAACUAAAAAUAUGUGUGAAAUAUUCUGGUUCUAGAGAGGC ......((((..---..))))(((.(((----((((...........(((((((.....)))))))..(((((.......))))).))))))).....))) ( -19.90, z-score = -0.74, R) >droPer1.super_14 937337 94 - 2168203 AACAAAGGCCAA---AAGGUCGCUGGGA----AUCAAUGGCACUUGAUGGUUACACAAAGUAACUAAAAAUAUGUGUGAAAUAUUCUGGUUCUAGAGAGGC ......((((..---..))))(((.(((----((((...((((....(((((((.....))))))).......))))((.....))))))))).....))) ( -21.10, z-score = -1.42, R) >consensus CAUAAAGGUGAA___AAGGUAGCCGAGCGAUCAUCAAUGCCACUUGAUGGUCACACAACGAGAUAAAAA__AUGUGUGAAAU_UUCAGGUUCCA_______ ......(((............))).................((((((...(((((((...............))))))).....))))))........... (-11.44 = -11.52 + 0.08)

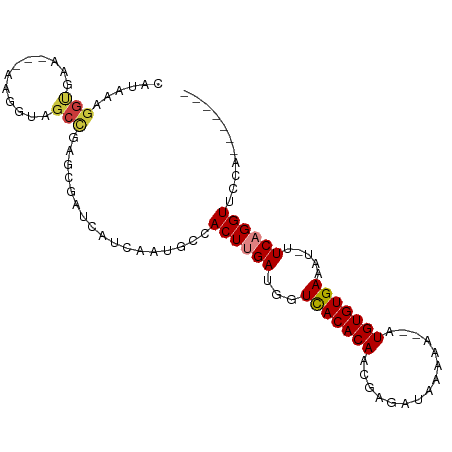

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:23:51 2011