
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 10,541,433 – 10,541,524 |
| Length | 91 |
| Max. P | 0.941690 |

| Location | 10,541,433 – 10,541,524 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 81.05 |
| Shannon entropy | 0.36413 |
| G+C content | 0.53249 |
| Mean single sequence MFE | -32.08 |
| Consensus MFE | -22.25 |
| Energy contribution | -23.17 |
| Covariance contribution | 0.92 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.47 |
| SVM RNA-class probability | 0.941690 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
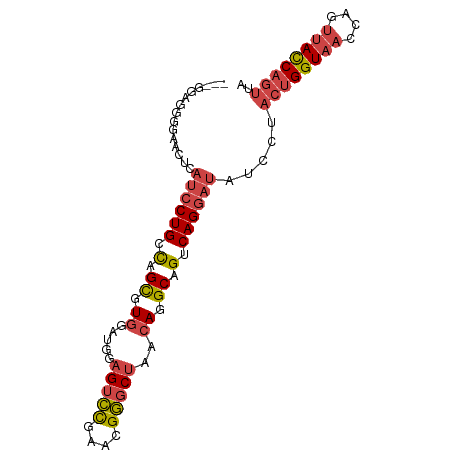
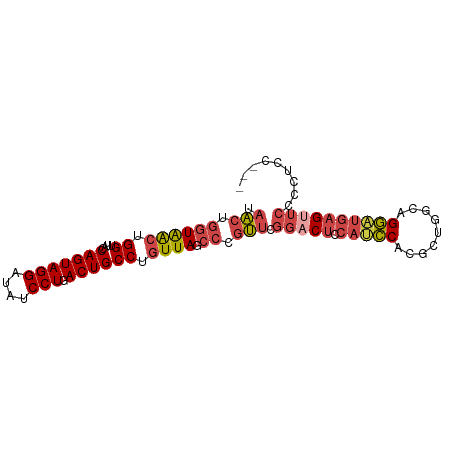
>dm3.chr2L 10541433 91 + 23011544 GGAGGAGGAGAACUCAUCCUGCCCGCGUGGAUGUAGUCCGAACGGGCUAACAGGCAGUCAGGAUAUCCUACUGGUAACCAGUUACCAGUUA (((.(((.....)))((((((.(.((.((....((((((....)))))).)).)).).)))))).))).((((((((....)))))))).. ( -38.20, z-score = -3.35, R) >droSim1.chr2L 10345449 91 + 22036055 GGAGGAGGGGAACUCAUCCUGCUAGCGUGGAUGGAGUCCGAACGGGCUAAAAGGCAGUCAGGAUAUCCUACUGGUAACCAGUUACCAGUUA (((.(((.....)))((((((((.((.(......(((((....)))))...).)))).)))))).))).((((((((....)))))))).. ( -34.30, z-score = -2.26, R) >droSec1.super_3 5968040 91 + 7220098 CGAGGAGGGGAACUCAUCCUGCCAGCGUGGAUGGAGUCCGAACGGGCUAACAGGCAGUCAGGAUUUCCUACUGGUAACCAGUUACCAGUUA .......(((((...((((((.(.((.((.....(((((....)))))..)).)).).)))))))))))((((((((....)))))))).. ( -34.50, z-score = -2.05, R) >droYak2.chr2L 6950298 88 + 22324452 ---GGACGAGAACUCAUCCUGCCCGCGUGGAUGGAGUCCAAACGGGCUAACAGGCAGUCAGGAUAUCCUACUGGUAACCAGUUACCAGUUA ---(((.((....))((((((.(.((.((.....(((((....)))))..)).)).).)))))).))).((((((((....)))))))).. ( -32.70, z-score = -2.46, R) >droEre2.scaffold_4929 11755208 88 - 26641161 ---GGACAGGACCUCAUCCUGCCAGUGUGGCUGCAGUCCGAACGGGCUAACAGGCAGUCAGGAUAUCCUACUGGUAACCAGUUACCAGUUA ---((.(((((.....))))))).....(((((((((((....))))).....))))))(((....)))((((((((....)))))))).. ( -35.80, z-score = -2.74, R) >droAna3.scaffold_12916 14391657 78 + 16180835 -----------CCUUUCACUGUCUGUGUGUGUGU-GUGUGUCCGU-CUAACAGGCAGUCAGGAUAUCCUACUGGUUACCAGCCAUCCUACA -----------......(((((((((........-..........-...))))))))).(((((......((((...))))..)))))... ( -17.00, z-score = 0.79, R) >consensus ___GGAGGGGAACUCAUCCUGCCAGCGUGGAUGGAGUCCGAACGGGCUAACAGGCAGUCAGGAUAUCCUACUGGUAACCAGUUACCAGUUA ...............((((((.(.((.((.....(((((....)))))..)).)).).)))))).....((((((((....)))))))).. (-22.25 = -23.17 + 0.92)


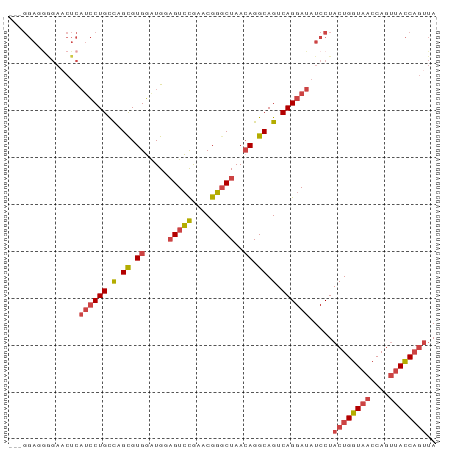
| Location | 10,541,433 – 10,541,524 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 91 |
| Reading direction | reverse |
| Mean pairwise identity | 81.05 |
| Shannon entropy | 0.36413 |
| G+C content | 0.53249 |
| Mean single sequence MFE | -28.89 |
| Consensus MFE | -17.43 |
| Energy contribution | -18.55 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.578873 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 10541433 91 - 23011544 UAACUGGUAACUGGUUACCAGUAGGAUAUCCUGACUGCCUGUUAGCCCGUUCGGACUACAUCCACGCGGGCAGGAUGAGUUCUCCUCCUCC ..((((((((....))))))))(((((((((((((.....))).((((((..(((.....)))..)))))))))))).....))))..... ( -33.80, z-score = -2.55, R) >droSim1.chr2L 10345449 91 - 22036055 UAACUGGUAACUGGUUACCAGUAGGAUAUCCUGACUGCCUUUUAGCCCGUUCGGACUCCAUCCACGCUAGCAGGAUGAGUUCCCCUCCUCC ..((((((((....)))))))).((((((((((.((((.....((.((....)).))........)).)))))))))...)))........ ( -24.82, z-score = -0.98, R) >droSec1.super_3 5968040 91 - 7220098 UAACUGGUAACUGGUUACCAGUAGGAAAUCCUGACUGCCUGUUAGCCCGUUCGGACUCCAUCCACGCUGGCAGGAUGAGUUCCCCUCCUCG ..((((((((....))))))))((((......((((.(((((((((..((..(((.....))))))))))))))...))))....)))).. ( -29.80, z-score = -1.93, R) >droYak2.chr2L 6950298 88 - 22324452 UAACUGGUAACUGGUUACCAGUAGGAUAUCCUGACUGCCUGUUAGCCCGUUUGGACUCCAUCCACGCGGGCAGGAUGAGUUCUCGUCC--- ((((.(((((((((...)))))(((....)))...)))).))))((((((.((((.....)))).)))))).((((((....))))))--- ( -37.80, z-score = -3.97, R) >droEre2.scaffold_4929 11755208 88 + 26641161 UAACUGGUAACUGGUUACCAGUAGGAUAUCCUGACUGCCUGUUAGCCCGUUCGGACUGCAGCCACACUGGCAGGAUGAGGUCCUGUCC--- ..((((((((....))))))))(((((((((((..((.((((.((.((....)).)))))).))......))))))...)))))....--- ( -30.00, z-score = -0.93, R) >droAna3.scaffold_12916 14391657 78 - 16180835 UGUAGGAUGGCUGGUAACCAGUAGGAUAUCCUGACUGCCUGUUAG-ACGGACACAC-ACACACACACAGACAGUGAAAGG----------- ..((((((((((((...)))))....)))))))((((.((((...-..(.......-.)......)))).))))......----------- ( -17.10, z-score = 0.49, R) >consensus UAACUGGUAACUGGUUACCAGUAGGAUAUCCUGACUGCCUGUUAGCCCGUUCGGACUCCAUCCACGCUGGCAGGAUGAGUUCCCCUCC___ .(((.((((((.((....(((((((....))).)))))).)))).)).))).(((((.(((((.........))))))))))......... (-17.43 = -18.55 + 1.12)

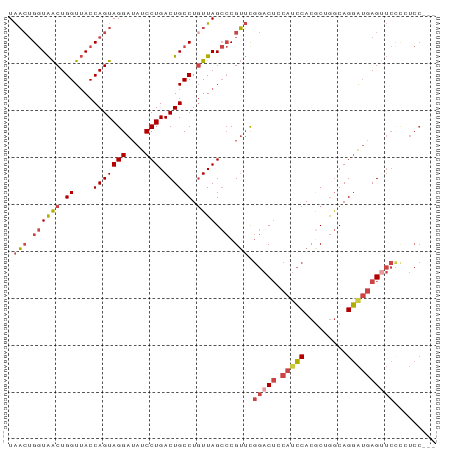
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:30:25 2011