
| Sequence ID | dm3.chrX |
|---|---|
| Location | 7,410,570 – 7,410,662 |
| Length | 92 |
| Max. P | 0.698915 |

| Location | 7,410,570 – 7,410,662 |
|---|---|
| Length | 92 |
| Sequences | 11 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 77.35 |
| Shannon entropy | 0.49113 |
| G+C content | 0.38357 |
| Mean single sequence MFE | -21.14 |
| Consensus MFE | -13.54 |
| Energy contribution | -13.42 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.05 |
| Mean z-score | -0.99 |
| Structure conservation index | 0.64 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.698915 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 7410570 92 - 22422827 ------UCCAGCUAGGCAGCGAACGACGAGCAACGGUCUCCGUUUUCAUUUGGUGUUUGAUUGGAUUUUCAAUUAUUCAUUAUUAUUGGUUGCAAUUU ------.........(((((.((..(.(((.(((((...))))))))...(((((..(((((((....)))))))..))))))..)).)))))..... ( -21.70, z-score = -1.23, R) >droSim1.chrX 5936474 89 - 17042790 ------UCCAGCCAGCCAGCGAACGACG---ACUGGUCUCCGUUUUCAUUUGGUGUUUGAUUGGAUUUUCAAUUAUUCAUUAUUAUUGGUUGCAAUUU ------..(((((((((((((.....))---.))))).............(((((..(((((((....)))))))..)))))....))))))...... ( -22.50, z-score = -1.87, R) >droSec1.super_24 618658 89 - 912625 ------UCCAGCCAGCCAGCGAACGACG---ACUGGUCUCCGUUUUCAUUUGGUGUUUGAUUGGAUUUUCAAUUAUUCAUUAUUAUUGGUUGCAAUUU ------..(((((((((((((.....))---.))))).............(((((..(((((((....)))))))..)))))....))))))...... ( -22.50, z-score = -1.87, R) >droEre2.scaffold_4690 16322433 81 - 18748788 --------------GCCAGUGAACGACG---ACUGGUCUCCGUUUUCAUUUGGUGUUUGAUUGGAUUUUCAAUUAUUCAUUAUUAUUGGUUGCAAUUU --------------((((((........---))))))....((..(((..(((((..(((((((....)))))))..)))))....)))..))..... ( -18.80, z-score = -1.66, R) >droAna3.scaffold_13117 261403 98 - 5790199 GCCAGUGGCUGUCAGAGCGAGAGCGAGUGAGUGCGGCCUCCGUUUUCAUUUGGUGUUUGAUUGGAUUUUCAAUUAUUCAUUAUUAUUGGUUGCAAUUU ((((((((((.((.....)).)))(((((((.((((...)))).)))))))((((..(((((((....)))))))..))))..)))))))........ ( -28.60, z-score = -1.73, R) >droPer1.super_14 846195 92 - 2168203 ---GAUGCUGAUGCUGAUGCUGCUGGC--CGUGU-GCCUCCGUUUUCAUUUGGUGUUUGAUUGGAUUUUCAAUUAUUCAUUAUUAUUGGUUGCAAUUU ---..(((....((..(((..(..(((--.....-)))..).........(((((..(((((((....)))))))..))))).)))..)).))).... ( -18.40, z-score = 0.33, R) >dp4.chrXL_group1e 11397765 92 - 12523060 ---GAUGCUGAUGCUGAUGCUGCUGGC--CGUGU-GCCUCCGUUUUCAUUUGGUGUUUGAUUGGAUUUUCAAUUAUUCAUUAUUAUUGGUUGCAAUUU ---..(((....((..(((..(..(((--.....-)))..).........(((((..(((((((....)))))))..))))).)))..)).))).... ( -18.40, z-score = 0.33, R) >droWil1.scaffold_180777 1381200 76 + 4753960 ---------------------GCUGGCAAUGUGC-GCCUCCGUUUUCAUUUGGUGUUUGAUUGGAUUUUCAAUUAUUCAUUAUUAUUGGUUGCAAUUU ---------------------(..(((.......-)))..)((..(((..(((((..(((((((....)))))))..)))))....)))..))..... ( -15.60, z-score = -0.71, R) >droVir3.scaffold_12970 3628226 94 - 11907090 ---UAUGGCAACGGCAAUGGCCAUGGCGUUCUGC-GCCUCCGUUUUCAUUUGGUGUUUGAUUGGAUUUUCAAUUAUUCAUUAUUAUUGGUUGCAAUUU ---....(((((((.(((((....((((.....)-))).))))).))...(((((..(((((((....)))))))..)))))......)))))..... ( -27.20, z-score = -1.96, R) >droMoj3.scaffold_6473 8128004 87 - 16943266 ---------UAUGGCAAAGGCAAUGGCAGCACG--GCCUCCGUUUUCAUUUGGUGUUUGAUUGGAUUUUCAAUUAUUCAUUAUUAUUGGUUGCAAUUU ---------...((...((((..((....))..--))))))((..(((..(((((..(((((((....)))))))..)))))....)))..))..... ( -19.10, z-score = -0.38, R) >droGri2.scaffold_15203 1735901 90 + 11997470 -------GCAAUGGCAAUGGCAAUAGCAUUGUGC-GCCUCCGUUUUCAUUUGGUGUUUGAUUGGAUUUUCAAUUAUUCAUUAUUAUUGGUUGCAAUUU -------(((((((.(((((.....((.....))-....))))).))...(((((..(((((((....)))))))..)))))......)))))..... ( -19.70, z-score = -0.12, R) >consensus _______CCAAUGAGAAAGCGAAUGGCG__GUGC_GCCUCCGUUUUCAUUUGGUGUUUGAUUGGAUUUUCAAUUAUUCAUUAUUAUUGGUUGCAAUUU ........................(((........)))...((..(((..(((((..(((((((....)))))))..)))))....)))..))..... (-13.54 = -13.42 + -0.12)

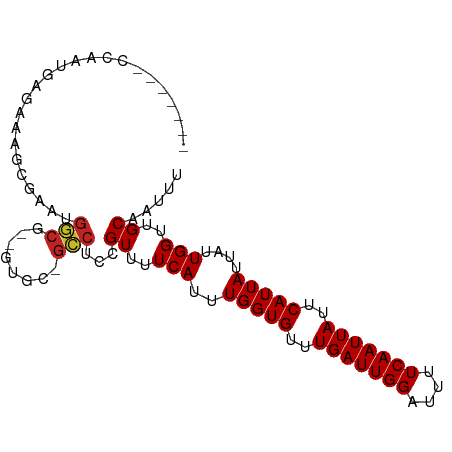

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:23:37 2011