
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 10,494,379 – 10,494,486 |
| Length | 107 |
| Max. P | 0.867072 |

| Location | 10,494,379 – 10,494,486 |
|---|---|
| Length | 107 |
| Sequences | 10 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 68.95 |
| Shannon entropy | 0.67567 |
| G+C content | 0.39381 |
| Mean single sequence MFE | -23.91 |
| Consensus MFE | -11.66 |
| Energy contribution | -11.41 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.40 |
| Mean z-score | -0.97 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.98 |
| SVM RNA-class probability | 0.867072 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 10494379 107 + 23011544 ACACGGCUAAUUAUGAUAU-AUCGCUAAUAAAAGAAUCAAGCAUAAU-ACCUAUGAAUAACUGCUGCAAUUUCUCGUUGCAAUAGUUGAUGCAGAACUGUUCAAAGCUG ...(((((.((((((....-....((......)).......))))))-.....((((((.((((((((((.....)))))).........))))...)))))).))))) ( -20.06, z-score = -0.59, R) >droSim1.chr2L 10297666 108 + 22036055 ACACGGAUCAUUACCAUAC-AUCGCUAAUUAAAGAAUCAGGCAUCAUUACCUAUGAAUAACUGCUGCAAUUUCUCGUUGCAAUAGUUGAUGCAGAACUGCUCAAAGCUG ....((.......))....-...(((............(((........))).(((.((((((.((((((.....)))))).))))))..(((....)))))).))).. ( -18.50, z-score = 0.19, R) >droSec1.super_3 5921731 108 + 7220098 ACACGGAUCAUUACGAUAU-AUCGCUAAUUAAAGAAUCAGGCAUCAUUACCUAUGAAUAACUGCUGCAAUUUCUCGUUGCAAUAGUUGAUGCAGAACUGCUCAAAGCUG ...(((.(((((((((...-.))).))))...........(((((((.....)))).((((((.((((((.....)))))).)))))).))).)).))).......... ( -19.00, z-score = 0.26, R) >droEre2.scaffold_4929 11707728 108 - 26641161 AAGAGGAUCAUUAUGAUAUUAUCCCUAAUAACCGUUUGCUUUAUCUU-ACCUAUGAAUAACUGCUGCAGUUUCUCGUUGCAGUAGUUGAUGCAGAACUGCUCAAAGCUA .((.((((.((......)).))))))...........(((((.....-..........(((((((((((.......)))))))))))((.(((....)))))))))).. ( -25.50, z-score = -1.85, R) >droYak2.chr2L 6901478 107 + 22324452 ACAAGGAUCAUUACAAUAU-AAAUCGAAUAACAAACUUCACUAUCAU-ACCUAUGAACAGCUGCUGCAGUUUCUCGUUGCAGUAGUUGAUGCAGAACUGCUCGAAGCUG ...................-...............((((....((((-....)))).((((((((((((.......))))))))))))..(((....)))..))))... ( -23.90, z-score = -1.20, R) >droAna3.scaffold_12916 14339035 107 + 16180835 CAAAGGAUAAGUACU-UGCCAUUUAGUAUGAACUAAUAUCCUCACUU-ACCAAUAAACAACUGCUGCAAUUUCUCAUUGCAAUAAUUAAUGCAAAACUGCUCAAAGCUG ...(((((((((.(.-(((......))).).)))..)))))).....-..............((((((((.....)))))..........(((....)))....))).. ( -15.40, z-score = -0.59, R) >droWil1.scaffold_180830 90487 109 + 119711 AAACGAAUCAUUGAAAUUUCAUUCCGGAAAGCGUUAAAUUUCCUACUGACCUAUAAACAACUGUUGCAGCUUCUCAUUGCAGUAGUUAAUGCAAAAUUGCUCAAAACUG ..........((((...........(((((........)))))...............(((((((((((.......)))))))))))...(((....)))))))..... ( -15.70, z-score = -0.07, R) >droVir3.scaffold_12963 8059420 104 + 20206255 ACCCAGUUAGAGAUA--CU--UUCCUGGGCCACUGCUUAGUUA-ACUCACCAAUGAACAGCUGCUGCAGCUUCUCGUUGCAGUAGUUAAUGCAGAACUGCUCAAAGCUG ...(((((.(((...--.(--(..((((((....))))))..)-)))).....(((..((((((((((((.....))))))))))))...(((....)))))).))))) ( -32.00, z-score = -1.72, R) >droMoj3.scaffold_6500 681722 107 + 32352404 ACGCUGUUAGGUUCUACUC--CUCCAGUGGCAUUAGACGCUUCCGCUCACCUAUGAACAGCUGCUGCAGCUUCUCGUUGCAGUAGUUGAUGCAGAACUGCUCAAAGCUA .(((((..(((.......)--)).)))))((....((.((....)))).....(((.(((((((((((((.....)))))))))))))..(((....))))))..)).. ( -35.80, z-score = -1.60, R) >droGri2.scaffold_15252 7495819 105 + 17193109 ACUUAGUUAAGGAGAUCUC--UUCUACAACUUUUAGAUGAGA--GCUCACCAAUGAACAACUGCUGCAGCUUUUCAUUGCAGUAGUUGAUGCAGAACUGCUCAAAGCUA ...(((((...(((.((((--.((((.......)))).))))--.))).....(((.((((((((((((.......))))))))))))..(((....)))))).))))) ( -33.20, z-score = -2.56, R) >consensus ACACGGAUAAUUACAAUAU_AUUCCUAAUAAAAUAAUCAAGUAUCAU_ACCUAUGAACAACUGCUGCAGUUUCUCGUUGCAGUAGUUGAUGCAGAACUGCUCAAAGCUG .........................................................((((((.((((((.....)))))).))))))..(((....)))......... (-11.66 = -11.41 + -0.25)


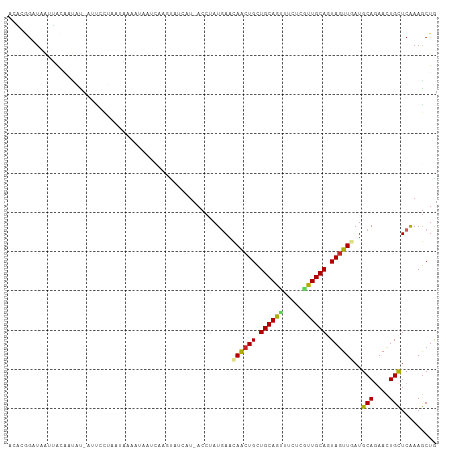
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:30:22 2011