
| Sequence ID | dm3.chrX |
|---|---|
| Location | 7,262,703 – 7,262,845 |
| Length | 142 |
| Max. P | 0.851494 |

| Location | 7,262,703 – 7,262,845 |
|---|---|
| Length | 142 |
| Sequences | 5 |
| Columns | 168 |
| Reading direction | reverse |
| Mean pairwise identity | 83.00 |
| Shannon entropy | 0.28332 |
| G+C content | 0.49032 |
| Mean single sequence MFE | -44.36 |
| Consensus MFE | -27.89 |
| Energy contribution | -28.21 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.851494 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
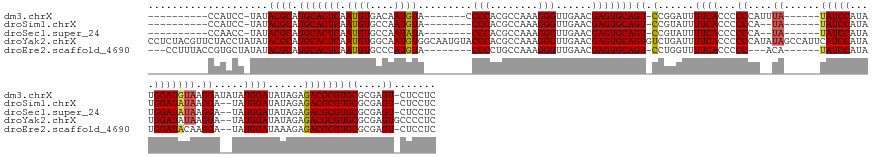
>dm3.chrX 7262703 142 - 22422827 ----------CCAUCC-UAUACGCAUGCACUCAAUGUGACAAUGUA-------CCCCACGCCAAAGGGUUGAACGAGUGCAGU-CCGGAUUUUCACCCCCCAUUUA------UAUCCAUAUGGAUGUAAGGAUAUAUGGAUAUAGAGACGCGUGCGCGAGU-CUCCUC ----------..((((-........(((((((...(((........-------...)))(((....))).....)))))))((-((((........)).....(((------(((((....))))))))))))....))))...(((((.((....)).))-)))... ( -41.10, z-score = -1.68, R) >droSim1.chrX 5799571 137 - 17042790 ----------CCAUCC-UAUACGCAUGCACUCAAUGUGCCAAUGUA--------CCCACGCCAAAGGGUUGAACGAGUGCAGU-CCGUAUUUUCACCCCCCA--UA------UAUCCAUAUGGAUAUAAGGA--UAUGGAUAUAGAGACGCGUGCGCGAGU-CUCCUC ----------..((((-.(((((..(((((((...((((....)))--------).((.(((....)))))...)))))))..-.))))).........((.--((------(((((....))))))).)).--...))))...(((((.((....)).))-)))... ( -43.60, z-score = -3.06, R) >droSec1.super_24 472237 137 - 912625 ----------CCAACC-UAUACGCAUGCACUCAAUGUGCCAAUAUA--------CCCACGCCAAAGGGUUGAACGAGUGCAGU-CCGUAUUUUCACCCCCCA--UA------UAUCCAUAUGGAUAUAAGGA--UAUGGAUAUAGAGACGCGUGCGCGAGU-CUCCUC ----------(((...-.(((((..(((((((.((((....))))(--------(((........)))).....)))))))..-.))))).........((.--((------(((((....))))))).)).--..))).....(((((.((....)).))-)))... ( -42.90, z-score = -3.33, R) >droYak2.chrX 7684020 166 + 21770863 CCUCUACGUUCUACCUAUAUACGCAUGCACUCAAUGUGGCAAUGUGGCAAUGUACCCACGCCAAAGGGUUGAACGAGUGCAGUGUCUGAUUUUCACCCCCCAUAUAGCCAUUCAUCCAUAUGGAUAUAAGGA--UAUGGAUAUAGAGACGCGUGCGCGAGUGCCCCUC .(((..(((((.((((......((((.......))))(((...((((........)))))))...)))).))))).(((((((((((.((.((((....((.((((.((((........)))).)))).)).--..)))).))..)))))).))))))))........ ( -49.30, z-score = -1.18, R) >droEre2.scaffold_4690 16170788 144 - 18748788 ---CCUUUACCGUGCUAUAUACGCAUGCACUCAAUGUGCCCAUGUA--------CCCCUGCCAAAGGGUUGAACGAGUGCAGU-CCUGGUUUUCACCCCC---ACA------UAUCCAUAUGGAUACAAGGA--UAUGGAUAAAGAGACGCGUGCGCGAGU-CUCCUC ---.....(((((((.....((((.(((((((...((((....)))--------)((((.....))))......)))))))((-((((((....)))...---.((------(((((...((....)).)))--)))).....)).)))))))))))).))-...... ( -44.90, z-score = -2.25, R) >consensus __________CCAUCC_UAUACGCAUGCACUCAAUGUGCCAAUGUA________CCCACGCCAAAGGGUUGAACGAGUGCAGU_CCGGAUUUUCACCCCCCA__UA______UAUCCAUAUGGAUAUAAGGA__UAUGGAUAUAGAGACGCGUGCGCGAGU_CUCCUC ....................((((.(((((((.((((....)))).........(((........)))......))))))).........((((.....(((..........(((((....)))))..........))).....)))).))))((....))....... (-27.89 = -28.21 + 0.32)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:23:13 2011