
| Sequence ID | dm3.chrX |
|---|---|
| Location | 7,195,552 – 7,195,663 |
| Length | 111 |
| Max. P | 0.638056 |

| Location | 7,195,552 – 7,195,663 |
|---|---|
| Length | 111 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 70.29 |
| Shannon entropy | 0.56419 |
| G+C content | 0.32199 |
| Mean single sequence MFE | -21.92 |
| Consensus MFE | -8.95 |
| Energy contribution | -8.90 |
| Covariance contribution | -0.05 |
| Combinations/Pair | 1.44 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.638056 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
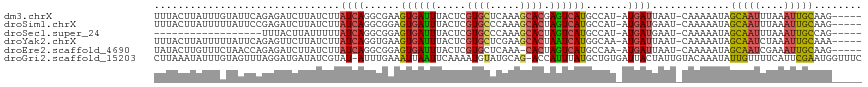
>dm3.chrX 7195552 111 - 22422827 UUUACUUAUUUGUAUUCAGAGAUCUUAUCUUAUCAGGCGAAGUGAUUUACUCGUGCUCAAAGCACGAGUCAUGCCAU-AUGAUUAAU-CAAAAAUAGCAAUUUAAAUUGCAAG----- ........((((...((((((((...)))))....((((((....)))((((((((.....))))))))...)))..-.))).....-))))....(((((....)))))...----- ( -25.80, z-score = -2.50, R) >droSim1.chrX 5746506 111 - 17042790 UUUACUUAUUUUUAUUCCGAGAUCUUAUCUUAUCAGGCGGAGUGAUUUACUCGUGCCCAAAGCACUAGUCAUGCCAU-AUGAUGAAU-CAAAAAUAGCAAUUUAAAUUGCAAG----- ......(((((((((((.(((((...)))))(((((((((((((...)))))((((.....))))......))))..-.))))))))-.)))))))(((((....)))))...----- ( -24.10, z-score = -2.09, R) >droSec1.super_24 404426 93 - 912625 ------------------UUUACUUAUUUUUAUCAGGCGGAGUGAUUUACUCGUGCCCAAAGCACUAGUCAUGCCAU-AUGAUGAAU-CAAAAAUAGCAAUUUAAAUUGCCAG----- ------------------..........((((((((((((((((...)))))((((.....))))......))))..-.))))))).-........(((((....)))))...----- ( -20.50, z-score = -2.02, R) >droYak2.chrX 7609555 111 + 21770863 UUUACUUAUUUUUAUUCAGAGUUCUUAUCUUAUCAGGUGAAGUGAUUUACUCGUGCUCGAAGCACUAAUCAUGGCAA-AUGAUUAAU-CAAAAAUAGCAAUCUAAAUUGCAAA----- ......(((((((.....((((....(((...((....))...)))..))))((((.....))))(((((((.....-)))))))..-.)))))))(((((....)))))...----- ( -22.10, z-score = -1.66, R) >droEre2.scaffold_4690 16099897 110 - 18748788 UAUACUUGUUUCUAACCAGAGAUCUUAUCUUAUCAGGCGGAGUGAUUUACUCGUGCUCAAA-CACUAGUCAUGCCAA-AUGAUUAAU-CAAAAAUAGCAAUCGAAAUUGCAAG----- ......(((((.......(((((...)))))....(((((((((...))))).)))).)))-)).(((((((.....-)))))))..-........(((((....)))))...----- ( -18.50, z-score = -0.30, R) >droGri2.scaffold_15203 287499 116 + 11997470 CUUAAAUAUUUGUAGUUUAGGAUGAUAUCGUAU-AUUUGAAAUUAAUUCAAAAUGUAUGCAG-ACCAUUUAUGCUGUGAUUACUAUUGUACAAAUAUUGUUUUCAUUCGAAUGGUUUC (((((((.......))))))).......(((((-(((((((.....))).))))))))).((-(((((((.....((((..((((((.....))))..))..))))..))))))))). ( -20.50, z-score = -0.60, R) >consensus UUUACUUAUUUGUAUUCAGAGAUCUUAUCUUAUCAGGCGAAGUGAUUUACUCGUGCUCAAAGCACUAGUCAUGCCAU_AUGAUUAAU_CAAAAAUAGCAAUUUAAAUUGCAAG_____ ...................................(((...((((((.....((((.....)))).))))))))).....................(((((....)))))........ ( -8.95 = -8.90 + -0.05)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:22:59 2011