
| Sequence ID | dm3.chrX |
|---|---|
| Location | 6,943,116 – 6,943,209 |
| Length | 93 |
| Max. P | 0.847346 |

| Location | 6,943,116 – 6,943,209 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 65.22 |
| Shannon entropy | 0.65064 |
| G+C content | 0.59222 |
| Mean single sequence MFE | -30.80 |
| Consensus MFE | -10.72 |
| Energy contribution | -11.90 |
| Covariance contribution | 1.18 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.35 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.747542 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
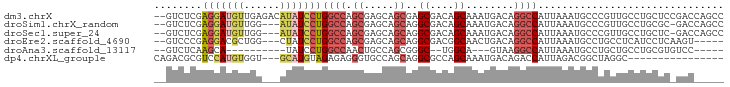
>dm3.chrX 6943116 93 + 22422827 --GUCUCGAGGAUGUUGAGACAUAUCCUGGCCAGCGAGCAGCGAGCGACAGCAAAUGACAGGCCAUUAAAUGCCCGUUGCCUGCUCCGACCAGCC --(((((((.....))))))).....((((...(.((((((...(((((.(((((((......))))...)))..))))))))))))..)))).. ( -31.60, z-score = -1.91, R) >droSim1.chrX_random 2203248 89 + 5698898 --GUCUCGAGGAUGUUGG---AUAUCCUGGCCAGCGAGCAGCAGGCGACAGCAAAUGACAGGCCAUUAAAUGCCCGUUGCCUGCGC-GACCAGCC --(((.(.(((((((...---))))))).).......((.(((((((((.(((((((......))))...)))..)))))))))))-)))..... ( -31.80, z-score = -1.18, R) >droSec1.super_24 163260 89 + 912625 --GUCUCGAGGAUGUUGG---AUAUCCUGGCCAGCGAGCAGCAGGCGACAGCAAAUGACAGGCCAUUAAAUGCCCGUUGCCUGCUC-GACCAGCC --......(((((((...---)))))))(((..(((((((...((((((.(((((((......))))...)))..)))))))))))-).)..))) ( -31.00, z-score = -1.13, R) >droEre2.scaffold_4690 15854480 85 + 18748788 --GUCCCGAGGACGCUGG---CUAUCCUGGCCAGCGAGCAGCAGGCGACGGCAACUGACAGGCCAUUAAAUGCCUGCCUCAUCCUCAAGU----- --.....(((((.(((((---((.....)))))))(((..(((((((..(((.........)))......)))))))))).)))))....----- ( -37.30, z-score = -2.94, R) >droAna3.scaffold_13117 3507788 73 - 5790199 --GUCUCAAGCA----------UAUCCUGGCCAACUGCCAGCGGGC--UGGCA---GUAAGGCCAUUAAAUGCCUGCUGCCUGCGUGUCC----- --.......(((----------(.(((((((.....))))).))((--.((((---(((.(((........)))))))))).))))))..----- ( -28.80, z-score = -1.64, R) >dp4.chrXL_group1e 5706711 76 - 12523060 CAGACGCGUCCAUGUGGU---GCAUGUAGAGAGGGUGCCAGCAGGCGCCAGCAAAUGACAGACCAUUAGACGGCUAGGC---------------- .((.(.((((...(((((---...(((...(..((((((....))))))..).....))).)))))..)))))))....---------------- ( -24.30, z-score = -0.90, R) >consensus __GUCUCGAGGAUGUUGG___AUAUCCUGGCCAGCGAGCAGCAGGCGACAGCAAAUGACAGGCCAUUAAAUGCCCGUUGCCUGCUC_GAC_____ ........(((((((......)))))))((((.((.....))..((....))........))))............................... (-10.72 = -11.90 + 1.18)



| Location | 6,943,116 – 6,943,209 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 65.22 |
| Shannon entropy | 0.65064 |
| G+C content | 0.59222 |
| Mean single sequence MFE | -33.53 |
| Consensus MFE | -11.43 |
| Energy contribution | -12.60 |
| Covariance contribution | 1.17 |
| Combinations/Pair | 1.45 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.34 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.90 |
| SVM RNA-class probability | 0.847346 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 6943116 93 - 22422827 GGCUGGUCGGAGCAGGCAACGGGCAUUUAAUGGCCUGUCAUUUGCUGUCGCUCGCUGCUCGCUGGCCAGGAUAUGUCUCAACAUCCUCGAGAC-- ..((((((((((((((..((((((........))))))..))))))...((.....))...)))))))).....(((((.........)))))-- ( -34.10, z-score = -1.06, R) >droSim1.chrX_random 2203248 89 - 5698898 GGCUGGUC-GCGCAGGCAACGGGCAUUUAAUGGCCUGUCAUUUGCUGUCGCCUGCUGCUCGCUGGCCAGGAUAU---CCAACAUCCUCGAGAC-- (((..((.-((((((((.((((.((...(((((....))))))))))).)))))).))..))..)))(((((..---.....)))))......-- ( -35.50, z-score = -1.68, R) >droSec1.super_24 163260 89 - 912625 GGCUGGUC-GAGCAGGCAACGGGCAUUUAAUGGCCUGUCAUUUGCUGUCGCCUGCUGCUCGCUGGCCAGGAUAU---CCAACAUCCUCGAGAC-- (((..(.(-((((((.((((((((........)))))).....((....)).))))))))))..)))(((((..---.....)))))......-- ( -35.50, z-score = -1.80, R) >droEre2.scaffold_4690 15854480 85 - 18748788 -----ACUUGAGGAUGAGGCAGGCAUUUAAUGGCCUGUCAGUUGCCGUCGCCUGCUGCUCGCUGGCCAGGAUAG---CCAGCGUCCUCGGGAC-- -----.((((((((((((((((((.....(((((.........))))).))))))..)))((((((.......)---))))))))))))))..-- ( -41.40, z-score = -2.90, R) >droAna3.scaffold_13117 3507788 73 + 5790199 -----GGACACGCAGGCAGCAGGCAUUUAAUGGCCUUAC---UGCCA--GCCCGCUGGCAGUUGGCCAGGAUA----------UGCUUGAGAC-- -----..............(((((((....(((((..((---(((((--(....)))))))).)))))....)----------))))))....-- ( -35.00, z-score = -3.73, R) >dp4.chrXL_group1e 5706711 76 + 12523060 ----------------GCCUAGCCGUCUAAUGGUCUGUCAUUUGCUGGCGCCUGCUGGCACCCUCUCUACAUGC---ACCACAUGGACGCGUCUG ----------------.....((((((((.((((((((....(((..((....))..)))........))).).---))))..)))))).))... ( -19.70, z-score = 0.12, R) >consensus _____GUC_GAGCAGGCAACAGGCAUUUAAUGGCCUGUCAUUUGCUGUCGCCUGCUGCUCGCUGGCCAGGAUAU___CCAACAUCCUCGAGAC__ ...........(((....((((((........))))))....))).((((((.((.....)).))).(((((..........)))))...))).. (-11.43 = -12.60 + 1.17)


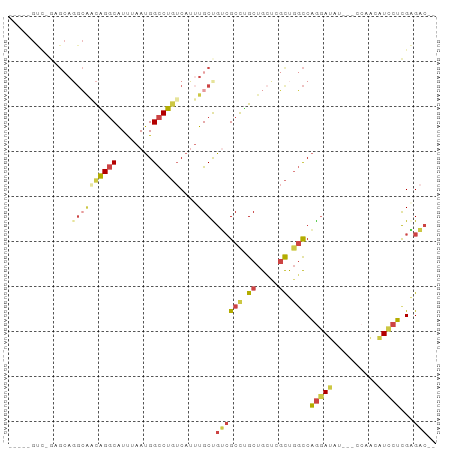
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:22:37 2011