
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 10,470,438 – 10,470,537 |
| Length | 99 |
| Max. P | 0.519600 |

| Location | 10,470,438 – 10,470,537 |
|---|---|
| Length | 99 |
| Sequences | 6 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 68.39 |
| Shannon entropy | 0.54521 |
| G+C content | 0.40981 |
| Mean single sequence MFE | -24.15 |
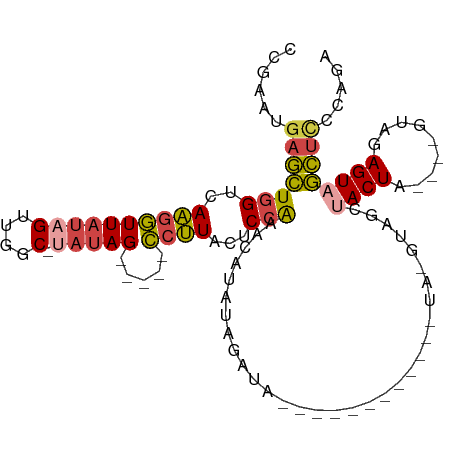
| Consensus MFE | -8.82 |
| Energy contribution | -8.91 |
| Covariance contribution | 0.09 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.519600 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
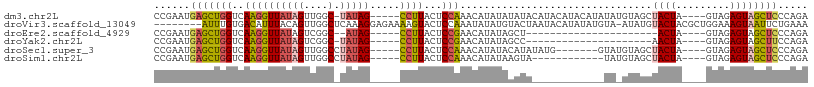
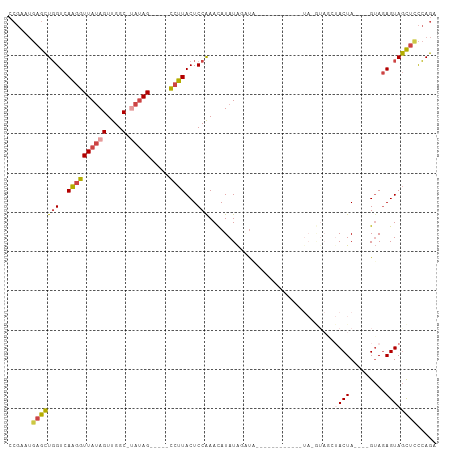
>dm3.chr2L 10470438 99 - 23011544 CCGAAUGAGCUGGUCAAGGUUAUAGUUGGC-UAUAG-----CCUUACUCCAAACAUAUAUAUACAUACAUACAUAUAUGUAGCUACUA----GUAGAGUAGCUCCCAGA .........((((..((((((((((....)-)))))-----)))).......((((((((.((......)).))))))))(((((((.----....))))))).)))). ( -30.00, z-score = -2.82, R) >droVir3.scaffold_13049 11289438 100 + 25233164 --------AUUUGUGACAUUUACAGUUGGCUCAAAGGAGAAAAGUACUCCAAAUAUAUGUACUAAUACAUAUAUGUA-AUAUGUACUACGCUGGAAAGUAAUUCUGAAA --------....(((((((((((............((((.......))))..(((((((((....))))))))))))-).))))..)))(((....))).......... ( -18.90, z-score = -0.91, R) >droEre2.scaffold_4929 11679315 76 + 26641161 CCGAAUGAGCUGGUCAAGGUUAUAGUCGGC--AUAG-----CCUUACUCCGAACAUAUAGCU----------------------ACUA----GUAGAGUAGCUCCCAGA .........((((..((((((((.(....)--))))-----)))).............((((----------------------(((.----....))))))).)))). ( -20.30, z-score = -0.97, R) >droYak2.chr2L 6876356 78 - 22324452 CCGAAUGAGCUGGUCAAGGUUAUAGUCGGC-UAUAG-----CCUUACUCCGAACAUAUAGCC---------------------AACUA----GUAGAGUAGCUUCCAGA ......((((((.((((((((((((....)-)))))-----))))............(((..---------------------..)))----...)).))))))..... ( -21.60, z-score = -1.52, R) >droSec1.super_3 5898396 93 - 7220098 CCGAAUGAGCUGGUCAAGGUUAUAGUUGGCCUAUAG-----CCUUACUCCAAACAUAUACAUAUAUG-------GUAUGUAGCUACUA----GUAGAGUAGCUCCCAGA .........((((..((((((((((.....))))))-----))))............(((((((...-------)))))))((((((.----....))))))..)))). ( -27.10, z-score = -1.59, R) >droSim1.chr2L 10264807 88 - 22036055 CCGAAUGAGCUGGUCAAGGUUAUAGUUGGCCUAUAG-----CCUUACUCCAAACAUAUAAGUA------------UAUGUAGCUACUA----GUAGAGUAGCUCCCAGA .........((((..((((((((((.....))))))-----)))).......((((((....)------------)))))(((((((.----....))))))).)))). ( -27.00, z-score = -2.47, R) >consensus CCGAAUGAGCUGGUCAAGGUUAUAGUUGGC_UAUAG_____CCUUACUCCAAACAUAUAGAUA____________UA_GUAGCUACUA____GUAGAGUAGCUCCCAGA ......((((((.((.....((((.((((...................))))...))))........................(((......))))).))))))..... ( -8.82 = -8.91 + 0.09)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:30:14 2011