
| Sequence ID | dm3.chrX |
|---|---|
| Location | 6,782,078 – 6,782,134 |
| Length | 56 |
| Max. P | 0.689654 |

| Location | 6,782,078 – 6,782,134 |
|---|---|
| Length | 56 |
| Sequences | 5 |
| Columns | 59 |
| Reading direction | forward |
| Mean pairwise identity | 80.88 |
| Shannon entropy | 0.33089 |
| G+C content | 0.46939 |
| Mean single sequence MFE | -10.92 |
| Consensus MFE | -8.66 |
| Energy contribution | -9.34 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.25 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.689654 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 6782078 56 + 22422827 UCGGCACACAACAAUUGAAUGGCCCCAA-GUAGACAGCCAUUU--UGCCGCCAUUCGCA .(((((..........(((((((.....-.......)))))))--)))))......... ( -12.20, z-score = -1.02, R) >droSim1.chrX 5311238 56 + 17042790 UCGGCACACAACAAUUGAAUGGCCCAAA-GUAGACAGCCAUUU--UGCCGCCAUCCACA .(((((..........(((((((.....-.......)))))))--)))))......... ( -12.20, z-score = -1.66, R) >droSec1.super_4 6165708 56 - 6179234 UCGGCACACAACAAUUGAAUGGCCCAAA-GUAGACAGCCAUUU--UGCCGCCAUCCACA .(((((..........(((((((.....-.......)))))))--)))))......... ( -12.20, z-score = -1.66, R) >droYak2.chrX 7197974 53 - 21770863 ----UCAAUAACAAUUGUAUGGCCCAAAAGUAGACAGCCAUUUCUUAUUGCCAUUCA-- ----.......((((...(((((.............))))).....)))).......-- ( -5.82, z-score = -0.16, R) >droEre2.scaffold_4690 15710007 51 + 18748788 UCGGCACAUAACAAUUGAAUGGCCCAAA-GUAGAUAGCCAUUU--UGCCGCCCA----- .(((((..........(((((((.....-.......)))))))--)))))....----- ( -12.20, z-score = -1.73, R) >consensus UCGGCACACAACAAUUGAAUGGCCCAAA_GUAGACAGCCAUUU__UGCCGCCAUCCACA .(((((..........(((((((.............)))))))..)))))......... ( -8.66 = -9.34 + 0.68)

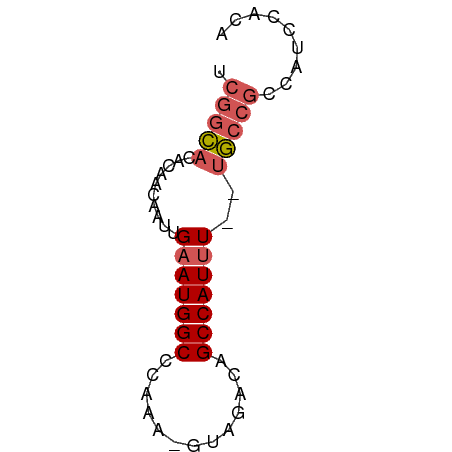

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:22:11 2011