
| Sequence ID | dm3.chrX |
|---|---|
| Location | 6,780,545 – 6,780,655 |
| Length | 110 |
| Max. P | 0.565599 |

| Location | 6,780,545 – 6,780,655 |
|---|---|
| Length | 110 |
| Sequences | 8 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 82.91 |
| Shannon entropy | 0.34308 |
| G+C content | 0.32674 |
| Mean single sequence MFE | -19.51 |
| Consensus MFE | -8.27 |
| Energy contribution | -9.11 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.565599 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
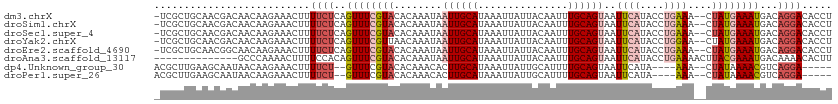
>dm3.chrX 6780545 110 + 22422827 -UCGCUGCAACGACAACAAGAAACUUUUCUCAGUUUCGUACACAAAUAAUUGCAUAAAUUAUUACAAUUUGCAGUAAUUCAUACCUGAAA--CUAUGAAAUGACAGGACACCU -..(((((((.........((((((......)))))).......(((((((.....))))))).....)))))))........((((...--...........))))...... ( -20.14, z-score = -2.52, R) >droSim1.chrX 5309764 110 + 17042790 -UCGCUGCAACGACAACAAGAAACUUUUCUCAGUUUCGUACACAAAUAAUUGCAUAAAUUAUUACAAUUUGCAGUAAUUCAUACCUGAAA--CUAUGAAAUGACAGGACACCU -..(((((((.........((((((......)))))).......(((((((.....))))))).....)))))))........((((...--...........))))...... ( -20.14, z-score = -2.52, R) >droSec1.super_4 6164227 110 - 6179234 -UCGCUGCAACGACAACAAGAAACUUUUCUCAGUUUCGUACACAAAUAAUUGCAUAAAUUAUUACAAUUUGCAGUAAUUCAUACCUGAAA--CUAUGAAAUGACAGGACACCU -..(((((((.........((((((......)))))).......(((((((.....))))))).....)))))))........((((...--...........))))...... ( -20.14, z-score = -2.52, R) >droYak2.chrX 7196335 110 - 21770863 -UCGCUGCAACGACAACAAGAAACUUUUCUCAGUUUCGUUAACAAAUAAUUGCAUAAAUUAUUACAAUUUGCAGUAAUUCAUACCUGGAA--CUAUGAAAUGACAGGACACCU -..(((((((.........((((((......)))))).......(((((((.....))))))).....)))))))........((((...--...........))))...... ( -21.44, z-score = -2.61, R) >droEre2.scaffold_4690 15708588 110 + 18748788 -UCGCUGCAACGGCAACAAGAAACUUUUCUCAGUUUCGUACACAAAUAAUUGCAUAAAUUAUUACAAUUUGCAGUAAUUCAUACCUGAAA--CUAUGAAAUGACAGGACACCU -..(((((((.(....)..((((((......)))))).......(((((((.....))))))).....)))))))........((((...--...........))))...... ( -21.04, z-score = -2.35, R) >droAna3.scaffold_13117 1619277 99 - 5790199 --------------GCCCAAAACUUUUCCACAGUUUCGUACACAAAUAAUUGCAUAAAUUAUUACAAUUUGCAGUAAUUCAUACCUGAAAACUUACGAAAUGACAAAACACUU --------------..................((((((((........((((((...............))))))..((((....))))....))))))))............ ( -12.56, z-score = -2.17, R) >dp4.Unknown_group_30 47024 100 - 91206 ACGCUUGAAGCAAUAACAAGAAACUUUUCU--GUUUCGUACACAAACACUUGCAUAAAUUAUUGCAUUUUGCAGUAAUUCAUA----AAA--CUAUAAAACGUCAGGA----- ...(((((.((((......(((((......--)))))((......))..))))...(((((((((.....)))))))))....----...--..........))))).----- ( -20.30, z-score = -2.56, R) >droPer1.super_26 685700 100 + 1349181 ACGCUUGAAGCAAUAACAAGAAACUUUUCU--GUUUCGUACACAAACACUUGCAUAAAUUAUUGCAUUUUGCAGUAAUUCAUA----AAA--CUAUAAAACGUCAGGA----- ...(((((.((((......(((((......--)))))((......))..))))...(((((((((.....)))))))))....----...--..........))))).----- ( -20.30, z-score = -2.56, R) >consensus _UCGCUGCAACGACAACAAGAAACUUUUCUCAGUUUCGUACACAAAUAAUUGCAUAAAUUAUUACAAUUUGCAGUAAUUCAUACCUGAAA__CUAUGAAAUGACAGGACACCU ..........................((((..((((((((........((((((...............))))))..(((......)))....))))))))...))))..... ( -8.27 = -9.11 + 0.84)

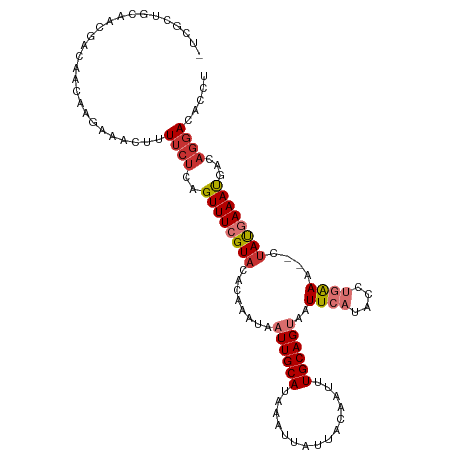

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:22:11 2011