
| Sequence ID | dm3.chrX |
|---|---|
| Location | 6,721,833 – 6,721,940 |
| Length | 107 |
| Max. P | 0.732072 |

| Location | 6,721,833 – 6,721,940 |
|---|---|
| Length | 107 |
| Sequences | 14 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 60.48 |
| Shannon entropy | 0.89785 |
| G+C content | 0.52536 |
| Mean single sequence MFE | -32.82 |
| Consensus MFE | -8.39 |
| Energy contribution | -8.01 |
| Covariance contribution | -0.39 |
| Combinations/Pair | 2.00 |
| Mean z-score | -1.41 |
| Structure conservation index | 0.26 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.732072 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 6721833 107 + 22422827 -----UUUUUUUGCACUUGCAGCAAAUUCUGGUUAACUCAAAGACGCAAGAUCCCACCU---ACAGUCAGGCCAUCGUUUACAUGGCCUGGUCAUUGGUCAGUUGCCAAGAAGCG -----....(((((.......)))))(((((((.((((....(((....).))..(((.---....(((((((((.(....)))))))))).....))).))))))).))))... ( -28.80, z-score = -1.39, R) >anoGam1.chrX 6782966 83 + 22145176 -----------------------------UCGCGGCGGGCAAGACGGGCGACCCGAUCU---UCGAGCACAUGCUGAUGUCGCUGGUGUGGGCGCUGGUGCGCAACUGGGAGUCG -----------------------------((.(((..(((...((.((((.((((....---.(.(((((((....)))).))).)..)))))))).)))).)..))).)).... ( -29.20, z-score = 1.41, R) >droVir3.scaffold_12928 3702846 102 - 7717345 ----------CUCCCUGCACAGCAAAUUCUGCUAACCCAAAAGUCGCAGGACUGCACAU---ACAGCCAGGCGAUCGUCUACUUGGCCUGGGCUUUCAUCAGCUGCAGGGAAUCG ----------.((((((((..(((..((((((.............)))))).)))....---....((((((.(.........).))))))(((......))))))))))).... ( -35.62, z-score = -2.44, R) >droMoj3.scaffold_6359 3920957 102 - 4525533 ----------CUCUUUUCACAGCAAAUCCUGCUCACCCAUAAGACGCAGGACUCCACGU---ACAGCCAGGCGAUAGUCUACCUGGCCUGGGCCUUUGUCAGCUGCCAGGAAUCG ----------.((((....((((...((((((((........)).))))))......(.---.((((((((..........))))).)))..)........))))..)))).... ( -29.10, z-score = -0.79, R) >droGri2.scaffold_15081 3723883 115 + 4274704 CUCUACCUCUCAUCUUUCGCAGGAAAUAUUACAGACGCCAAAGUCCCAGGACUGCUCAUUGCACAGCCAGGCGAUAGUUUAUUUAGCCUGGGCGCACAUCCAUUGCGGUGAAUCG ............((..((((((((.........((.((...((((....))))))))..(((.(..((((((.............))))))).)))..)))..))))).)).... ( -28.02, z-score = -0.19, R) >droWil1.scaffold_180777 2834663 109 - 4753960 ---UCUCUUUGUCCACUUACAGCAAAUACUUGUGACACAGAAUCCUCAAGAUUGCUCCU---UCAGCCAGGCUAUUGUCUAUUUGGCCUGGGCAUGGAUAUGCUGCCAAGAGGCC ---......((((((.....(((((...((((.((.......))..)))).)))))...---....((((((((.........))))))))...))))))....(((....))). ( -28.60, z-score = 0.06, R) >droPer1.super_15 1100617 110 + 2181545 --UUAUUCCAUUCCUCCUGCAGCAAAUCCUCGUCACACAAAAGACGCAGGAUGCCACCU---ACAGCCAGGCGCUGGUAUACCUCGCCUGGGCCUGGAUCAGCUGCCAGGAGGCG --..........(((((((((((..(((((((((........)))).))))).......---.((((((((((..((....)).)))))))..))).....)))).))))))).. ( -49.10, z-score = -4.83, R) >dp4.chrXL_group1a 8705056 110 + 9151740 --UUAUUCCAUUCCUCCUGCAGCAAAUCCUCGUCACACAAAAGACGCAGGAUGCCACCU---ACAGCCAGGCGCUGGUGUACCUCGCCUGGGCCUGGAUCAGCUGCCAGGAGGCG --..........(((((((((((..(((((((((........)))).))))).......---....((((((.(.((((.....)))).).))))))....)))).))))))).. ( -51.10, z-score = -4.92, R) >droAna3.scaffold_13117 521810 106 + 5790199 ----UAAUUUGUUUCCUUAGA--AAAUCUUGCUCACCCAGAAGGCCCAGGACGCCACCU---ACAGCCAGGCAAUCGUGUACCUGGCCUGGGCAUGGGUCUGCUGCCAGGAGUCU ----........(((((.((.--....)).((..(((((....(((((((....)....---...((((((..........)))))))))))).)))))..))....)))))... ( -33.50, z-score = 0.13, R) >droEre2.scaffold_4690 15649778 108 + 18748788 ----UUUGUCACUUCCUUGCAGCAAAUUCUGCUCACCCAAAAGACGCCAGAUGCCACCU---ACAGUCAGGCUAUCGUUUACAUGGCCUGGGCUUUGGUUAGUUGCCAAGAAGCA ----.......((((...(((((....(((...........))).((((((.(((((..---...))((((((((.(....))))))))))))))))))..)))))...)))).. ( -30.10, z-score = -0.58, R) >droYak2.chrX 2689316 108 + 21770863 ----UUUUUCACUUCCUUGCAGCAAAUUCUGCUCACCCAAAAGACGCAAGAUGCCACCU---ACAGUCAGGCUAUCGUUUACUUGGCCUGGGCUUGGGUCAGUUGCCAAGAAGCA ----.......((((...(((((..((..(((((........)).)))..))...((((---(.((((((((((.........)))))).)))))))))..)))))...)))).. ( -30.70, z-score = -0.69, R) >droSec1.super_4 6104958 109 - 6179234 ---UUUUGUCACUUCCUUGCAGCAAAUUCUGGUUAACCCACAGACGCAAGAUCCCACCU---ACAGUCAGGCCAUCGUUUACAUGGCCUGGGCCUUGAUCAGUUGCCAAGAAUCG ---............((((..((((...(((((((((((...(((...((.......))---...)))(((((((.(....)))))))))))..))))))))))))))))..... ( -30.10, z-score = -1.99, R) >droSim1.chrX 5276763 109 + 17042790 ---UUUUGUCUCUUCCUUGCAGCAAAUUCUGGUUAACCCACAGACGCAAGAUCCCACCU---ACAGUCAGGCCAUCGUUUACAUGGCCUGGGCCUUGAUCAGUUGCCAAGAAUCG ---............((((..((((...(((((((((((...(((...((.......))---...)))(((((((.(....)))))))))))..))))))))))))))))..... ( -30.10, z-score = -1.96, R) >triCas2.ChLG3 30418812 95 - 32080666 -----------------UUGGCGAUAUGGUAACGUCUGGAAGAACGUCCGGUUCUGAAU---UCGAACGAAUUGUAAUUUUUAUUACACUGGCGUUGCUACAAAUGCGGGAGACU -----------------...(((...((((((((((.(..(((((.....))))).(((---((....)))))(((((....))))).).))))))))))....)))((....)) ( -25.40, z-score = -1.52, R) >consensus _____UUUU_UUUCCCUUGCAGCAAAUUCUGGUCACCCAAAAGACGCAAGAUGCCACCU___ACAGCCAGGCGAUCGUUUACAUGGCCUGGGCCUUGAUCAGCUGCCAGGAAUCG ..................................................................((((((.............))))))........................ ( -8.39 = -8.01 + -0.39)


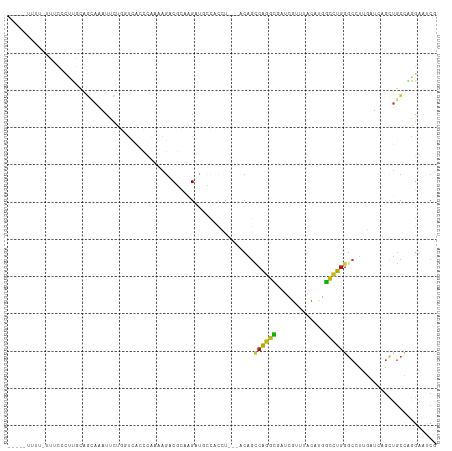
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:22:00 2011