| Sequence ID | dm3.chrX |
|---|---|
| Location | 6,514,022 – 6,514,122 |
| Length | 100 |
| Max. P | 0.593201 |

| Location | 6,514,022 – 6,514,122 |
|---|---|
| Length | 100 |
| Sequences | 8 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 73.01 |
| Shannon entropy | 0.50196 |
| G+C content | 0.41875 |
| Mean single sequence MFE | -20.96 |
| Consensus MFE | -6.33 |
| Energy contribution | -7.19 |
| Covariance contribution | 0.86 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.30 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.593201 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
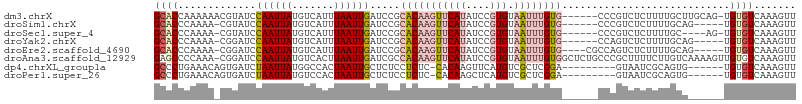
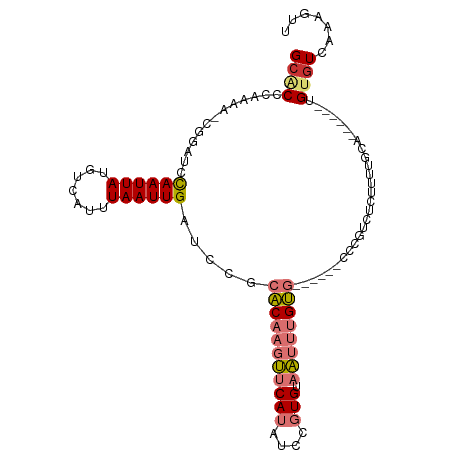
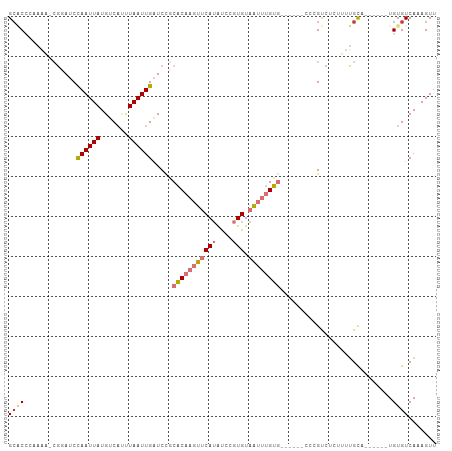
>dm3.chrX 6514022 100 + 22422827 GCACCAAAAAACGUAUCCAAUUAUGUCAUUUAAUUGAUCCGCACAAGUUCAUAUCCGUGUAAUUUGUG------CCCGUCUCUUUUGCUUGCAG-UGUGUCAAAGUU (((.(((((.(((....((((((.......))))))....((((((((((((....))).))))))))------).)))...)))))..)))..-............ ( -20.20, z-score = -2.54, R) >droSim1.chrX 5118610 95 + 17042790 GCACCCAAAA-CGUAUCCAAUUAUGUCAUUUAAUUGAUCCGCACAAGUUCAUAUCCGUGUAAUUUGUG------CCCGUCUCUUUUGCAG-----UGUGUCAAAGUU ((((.(((((-((((......))))).........(((..((((((((((((....))).))))))))------)..)))...))))..)-----)))......... ( -19.70, z-score = -3.14, R) >droSec1.super_4 5888677 95 - 6179234 GCACCCAAAA-CGUAUCCAAUUAUGUCAUUUAAUUGAUCCGCACAAGUUCAUAUCCGUGUAAUUUGUG------CCCGUCUCUUUUGC----AG-UGUGUCAAAGUU ((((.(((((-((((......))))).........(((..((((((((((((....))).))))))))------)..)))...)))).----.)-)))......... ( -19.70, z-score = -3.14, R) >droYak2.chrX 2477291 95 + 21770863 GCACCCAAAA-CGGAUCCAAUUAUGUCAUUUAAUUGAUCCGCACAAGUUCAUAUCCGUGUAAUUUGUG------CCAGUCUCUUUUGCAG-----UGUGUCAAAGUU ((((.(((((-.(((((.(((((.......))))))))))((((((((((((....))).))))))))------).......)))))..)-----)))......... ( -25.50, z-score = -4.19, R) >droEre2.scaffold_4690 15441376 97 + 18748788 GCACCCAAAA-CGGAUCCAAUUAUGUCAUUUAAUUGAUCCGCACAAGUUCAUAUCCGUGUAAUUUGUG----CGCCAGUCUCUUUUGCAG-----UGUGUCAAAGUU ((((.(((((-.(((((.(((((.......))))))))))((((((((((((....))).))))))))----).........)))))..)-----)))......... ( -26.40, z-score = -4.02, R) >droAna3.scaffold_12929 3083365 106 + 3277472 GAGCCCCAAA-CGGAUCCAAUUAUGUCACUUAAUUGAUCGCCACAAGUUCAUAUCCGUGUAAUUUGUGGCUCUGCCCGCUUUUCUUGUCAAAAGUUGUGUCAAAGUU ((((......-..((((.(((((.......)))))))))(((((((((((((....))).)))))))))).......))))...(((.((.......)).))).... ( -24.00, z-score = -2.17, R) >dp4.chrXL_group1a 315352 91 + 9151740 GCCCUGAAACAGUGAUCUAAUUAUGGCCACUAAUUGCUCUCCUCUC-CACAAGUUCAUGUCGCUCGGA---------GUAAUCGCAGUG------UGUGUCAAAGUU ....(((.((((((..(.......)..)))).(((((......(((-(...(((.......))).)))---------).....))))).------.)).)))..... ( -15.90, z-score = 0.43, R) >droPer1.super_26 997358 91 - 1349181 GCCCUGAAACAGUGAUCUAAUUAUGUCCACUAAUUGCUCUCCUCUC-CACAAGCUCAUGUCGCUCGGA---------GUAAUCGCAGUG------UGUGUCAAAGUU ....(((.((((((..(.......)..)))).(((((......(((-(...(((.......))).)))---------).....))))).------.)).)))..... ( -16.30, z-score = -0.26, R) >consensus GCACCCAAAA_CGGAUCCAAUUAUGUCAUUUAAUUGAUCCGCACAAGUUCAUAUCCGUGUAAUUUGUG______CCCGUCUCUUUUGCA______UGUGUCAAAGUU ((((.............((((((.......)))))).....(((((((((((....))).))))))))............................))))....... ( -6.33 = -7.19 + 0.86)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:21:20 2011