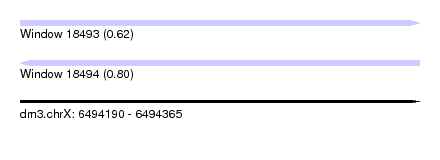

| Sequence ID | dm3.chrX |
|---|---|
| Location | 6,494,190 – 6,494,365 |
| Length | 175 |
| Max. P | 0.795255 |

| Location | 6,494,190 – 6,494,365 |
|---|---|
| Length | 175 |
| Sequences | 4 |
| Columns | 176 |
| Reading direction | forward |
| Mean pairwise identity | 96.01 |
| Shannon entropy | 0.06314 |
| G+C content | 0.35478 |
| Mean single sequence MFE | -43.50 |
| Consensus MFE | -34.24 |
| Energy contribution | -33.55 |
| Covariance contribution | -0.69 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.619825 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
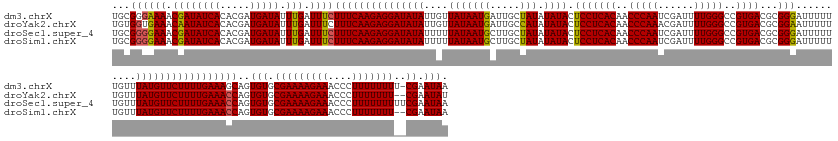
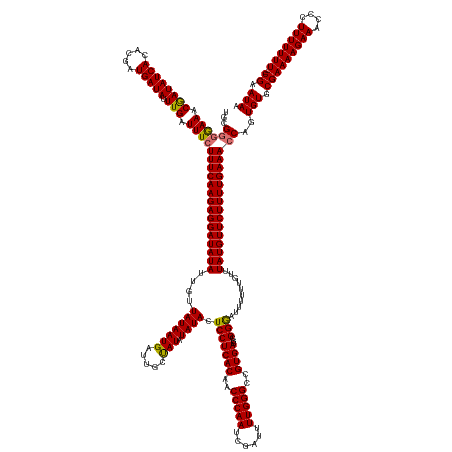
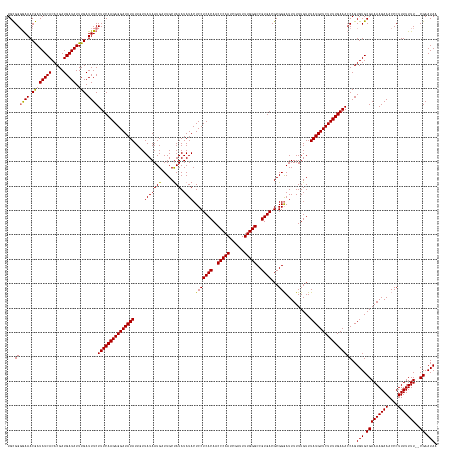
>dm3.chrX 6494190 175 + 22422827 UGCGGGAAAACGAUAUCACACGAUGAUAUUUGAUUUCUUUCAAGAGGAUAUAUUGUUAUAAUGAUUGCUAUAUAUACUCCUCACAACCCAAUCGAUUUUGGGCCGUGACGCGGGAUUUUUUGUUUAUGUUCUUUUGAAAGCAGUGUGCGAAAAGAAACCCUUUUUUUU-CGAAUAA (((.(((((.((((((((.....))))).))).)))))((((((((((((((....((((........))))....((((((((..(((((......)))))..)))).).)))..........)))))))))))))).))).(((.(((((((((.....)))))))-)).))). ( -43.00, z-score = -1.83, R) >droYak2.chrX 2460946 174 + 21770863 UGUGGUGAAACAAUAUCACACGAUGAUAUUUGAUUUCUUUCAAGAGGAUAUAUUGUUAUAAUGAUUGCCAUAUAUACUCCUCACAACCCAAACGAUUUUGGGCCGUGACGCGGAAUUUUUUGUUUAUGUUCUUUUGAAACCAGUGUGCGAAAAGAAACCCUUUUUUU--CGAAUAU ..((((((((((((((((.....))))).))).)))).((((((((((((((....(((((((.....))).)))).(((((((..((((((....))))))..))))...)))..........))))))))))))))))))((((.(((((((((....)))))))--)).)))) ( -44.10, z-score = -2.83, R) >droSec1.super_4 5872074 176 - 6179234 UGCGGGGAAACGAUAUCACACGAUGAUAUUUGAUUUCUUUCAAGAGGAUAUAUUUUUAUAAUGCUUGCUAUAUAUACUCCUCACAACCCAAUCGAUUUUGGGCCGUGACGCGGGAUUUUUUGUUUAUGUUCUUUUGAAACCAGUGUGCGAAAAGAAACCCUUUUUUUUUCGAAUAA ...(((((((((((((((.....))))).))).)))))((((((((((((((.......(((.(((((............((((..(((((......)))))..)))).)))))))).......)))))))))))))).))..(((.((((((((((....)))))))))).))). ( -43.76, z-score = -2.16, R) >droSim1.chrX 5101464 174 + 17042790 UGCGGGGAAACGAUAUCACACGAUGAUAUUUGAUUUCUUUCAAGAGGAUAUAUUUUUAUAAUGCUUGCUAUAUAUACUCCUCACAACCCAAUCGAUUUUGGGCCGUGACGCGGGAUUUUUUGUUUAUGUUCUUUUGAAACCAGUGUGCGAAAAGAAACCCUUUUUUU--CGAAUAA ...(((((((((((((((.....))))).))).)))))((((((((((((((.......(((.(((((............((((..(((((......)))))..)))).)))))))).......)))))))))))))).))..(((.(((((((((....)))))))--)).))). ( -43.16, z-score = -2.03, R) >consensus UGCGGGGAAACGAUAUCACACGAUGAUAUUUGAUUUCUUUCAAGAGGAUAUAUUGUUAUAAUGAUUGCUAUAUAUACUCCUCACAACCCAAUCGAUUUUGGGCCGUGACGCGGGAUUUUUUGUUUAUGUUCUUUUGAAACCAGUGUGCGAAAAGAAACCCUUUUUUU__CGAAUAA .((((((((.((((((((.....))))).))).)))))((((((((((((((....(((((((.....))).)))).(((((((..(((((......)))))..))))...)))..........)))))))))))))).......)))((((((.....))))))........... (-34.24 = -33.55 + -0.69)
| Location | 6,494,190 – 6,494,365 |
|---|---|
| Length | 175 |
| Sequences | 4 |
| Columns | 176 |
| Reading direction | reverse |
| Mean pairwise identity | 96.01 |
| Shannon entropy | 0.06314 |
| G+C content | 0.35478 |
| Mean single sequence MFE | -37.31 |
| Consensus MFE | -35.17 |
| Energy contribution | -35.66 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.94 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.795255 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
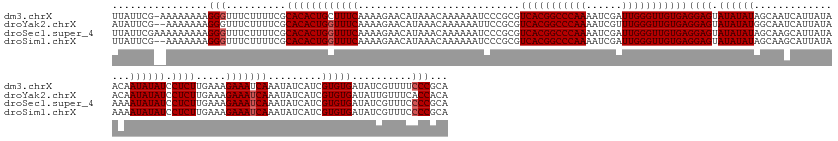
>dm3.chrX 6494190 175 - 22422827 UUAUUCG-AAAAAAAAGGGUUUCUUUUCGCACACUGCUUUCAAAAGAACAUAAACAAAAAAUCCCGCGUCACGGCCCAAAAUCGAUUGGGUUGUGAGGAGUAUAUAUAGCAAUCAUUAUAACAAUAUAUCCUCUUGAAAGAAAUCAAAUAUCAUCGUGUGAUAUCGUUUUCCCGCA ......(-((((.....((((((((((.((.....))...............................(((((((((((......)))))))))))((((.((((((................)))))).)))).))))))))))..(((((((...)))))))..)))))..... ( -35.99, z-score = -1.42, R) >droYak2.chrX 2460946 174 - 21770863 AUAUUCG--AAAAAAAGGGUUUCUUUUCGCACACUGGUUUCAAAAGAACAUAAACAAAAAAUUCCGCGUCACGGCCCAAAAUCGUUUGGGUUGUGAGGAGUAUAUAUGGCAAUCAUUAUAACAAUAUAUCCUCUUGAAAGAAAUCAAAUAUCAUCGUGUGAUAUUGUUUCACCACA ......(--(((.....(((((((((((((((....))(((....))).................)))((((((((((((....))))))))))))((((.((((((................)))))).)))).)))))))))).((((((((...)))))))).))))...... ( -39.19, z-score = -2.01, R) >droSec1.super_4 5872074 176 + 6179234 UUAUUCGAAAAAAAAAGGGUUUCUUUUCGCACACUGGUUUCAAAAGAACAUAAACAAAAAAUCCCGCGUCACGGCCCAAAAUCGAUUGGGUUGUGAGGAGUAUAUAUAGCAAGCAUUAUAAAAAUAUAUCCUCUUGAAAGAAAUCAAAUAUCAUCGUGUGAUAUCGUUUCCCCGCA ................(((..........((((((((((((...........................(((((((((((......)))))))))))((((.((((((................)))))).)))).....))))))).........)))))..........)))... ( -37.04, z-score = -1.39, R) >droSim1.chrX 5101464 174 - 17042790 UUAUUCG--AAAAAAAGGGUUUCUUUUCGCACACUGGUUUCAAAAGAACAUAAACAAAAAAUCCCGCGUCACGGCCCAAAAUCGAUUGGGUUGUGAGGAGUAUAUAUAGCAAGCAUUAUAAAAAUAUAUCCUCUUGAAAGAAAUCAAAUAUCAUCGUGUGAUAUCGUUUCCCCGCA .......--.......(((..........((((((((((((...........................(((((((((((......)))))))))))((((.((((((................)))))).)))).....))))))).........)))))..........)))... ( -37.04, z-score = -1.37, R) >consensus UUAUUCG__AAAAAAAGGGUUUCUUUUCGCACACUGGUUUCAAAAGAACAUAAACAAAAAAUCCCGCGUCACGGCCCAAAAUCGAUUGGGUUGUGAGGAGUAUAUAUAGCAAGCAUUAUAAAAAUAUAUCCUCUUGAAAGAAAUCAAAUAUCAUCGUGUGAUAUCGUUUCCCCGCA ................(((..........((((((((((((...........................(((((((((((......)))))))))))((((.((((((................)))))).)))).....))))))).........)))))..........)))... (-35.17 = -35.66 + 0.50)
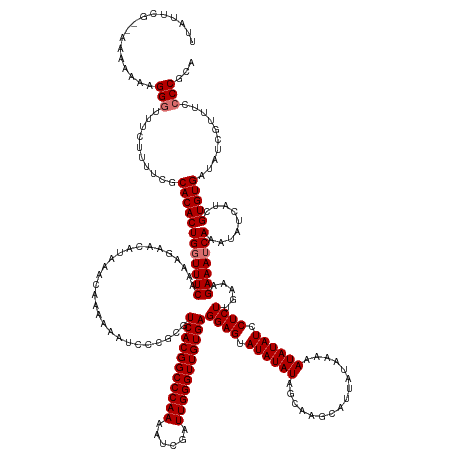
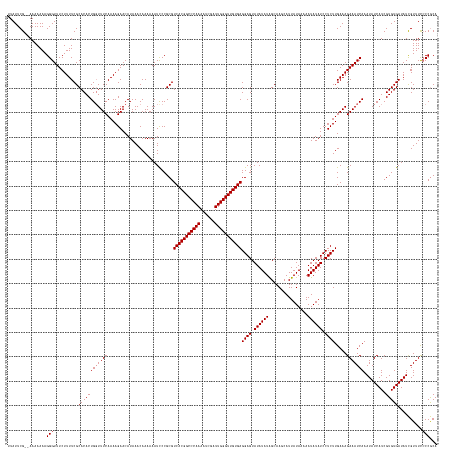
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:21:15 2011