
| Sequence ID | dm3.chrX |
|---|---|
| Location | 6,486,188 – 6,486,286 |
| Length | 98 |
| Max. P | 0.982207 |

| Location | 6,486,188 – 6,486,286 |
|---|---|
| Length | 98 |
| Sequences | 3 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 84.78 |
| Shannon entropy | 0.19678 |
| G+C content | 0.47953 |
| Mean single sequence MFE | -33.27 |
| Consensus MFE | -21.20 |
| Energy contribution | -21.53 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.37 |
| Structure conservation index | 0.64 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.10 |
| SVM RNA-class probability | 0.982207 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
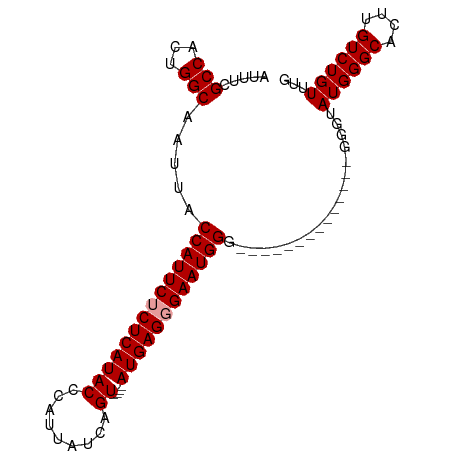
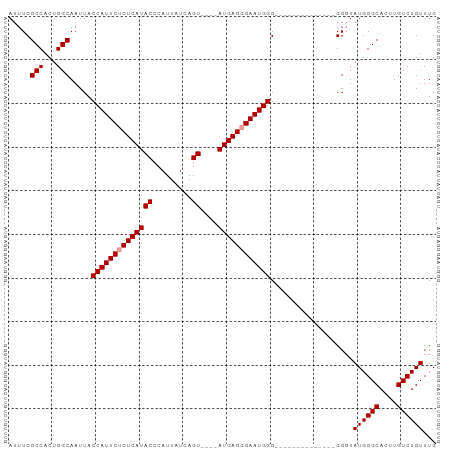
>dm3.chrX 6486188 98 + 22422827 AUUUCGCCACUGGCAAUUACCAUUCUCUCAUACCCAUUAUCAGUGAGUAUGAGUGAAUGGGUGGCGGGGGCGGGGGGGUAUGGGCACUUGUCUGUUUG .(..((((.(((.((....((((((.(((((((.(((.....))).))))))).)))))).)).))).))))..)....((((((....))))))... ( -41.40, z-score = -3.65, R) >droSim1.chrX 5093526 80 + 17042790 AUUUCGCCACUGGCAAUUACCAUUCUCUCAUACCCAUUAUCAGU----AUGAGGGAAUGGG--------------GGGUAUGGGCAGUUGUCUGUUUG ((..((((...))).....((((((((((((((.........))----)))))))))))))--------------..))..((((((....)))))). ( -30.20, z-score = -3.30, R) >droSec1.super_4 5864275 79 - 6179234 AUUUCGCCACUGGCAAUUACCAUUCUCUCAUACCCAUUAUCAGU----AUGAGGGAAUGGG--------------GG-UAUGGGCACUUGUCUGUUUG .....(((...))).((..((((((((((((((.........))----)))))))))))).--------------.)-)((((((....))))))... ( -28.20, z-score = -3.17, R) >consensus AUUUCGCCACUGGCAAUUACCAUUCUCUCAUACCCAUUAUCAGU____AUGAGGGAAUGGG______________GGGUAUGGGCACUUGUCUGUUUG .....(((...))).....((((((((((((....((.....))....))))))))))))...................((((((....))))))... (-21.20 = -21.53 + 0.33)

| Location | 6,486,188 – 6,486,286 |
|---|---|
| Length | 98 |
| Sequences | 3 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 84.78 |
| Shannon entropy | 0.19678 |
| G+C content | 0.47953 |
| Mean single sequence MFE | -25.30 |
| Consensus MFE | -16.93 |
| Energy contribution | -16.93 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.14 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.91 |
| SVM RNA-class probability | 0.974721 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
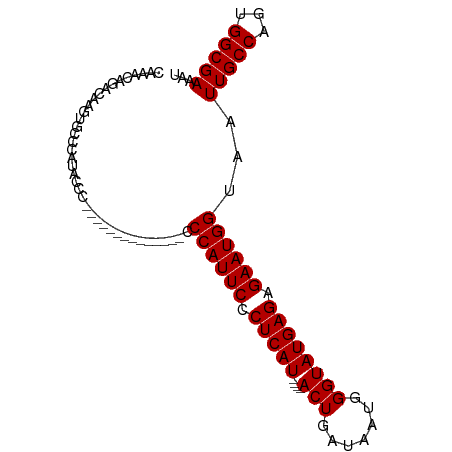
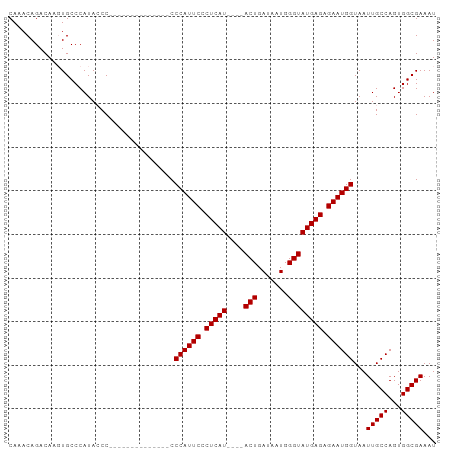
>dm3.chrX 6486188 98 - 22422827 CAAACAGACAAGUGCCCAUACCCCCCCGCCCCCGCCACCCAUUCACUCAUACUCACUGAUAAUGGGUAUGAGAGAAUGGUAAUUGCCAGUGGCGAAAU ................................((((((((((((.((((((((((.......)))))))))).)))))).........)))))).... ( -30.50, z-score = -4.05, R) >droSim1.chrX 5093526 80 - 17042790 CAAACAGACAACUGCCCAUACCC--------------CCCAUUCCCUCAU----ACUGAUAAUGGGUAUGAGAGAAUGGUAAUUGCCAGUGGCGAAAU ....(((....))).........--------------.((((((.(((((----(((.......)))))))).))))))...(((((...)))))... ( -23.00, z-score = -2.83, R) >droSec1.super_4 5864275 79 + 6179234 CAAACAGACAAGUGCCCAUA-CC--------------CCCAUUCCCUCAU----ACUGAUAAUGGGUAUGAGAGAAUGGUAAUUGCCAGUGGCGAAAU ....................-..--------------.((((((.(((((----(((.......)))))))).))))))...(((((...)))))... ( -22.40, z-score = -2.55, R) >consensus CAAACAGACAAGUGCCCAUACCC______________CCCAUUCCCUCAU____ACUGAUAAUGGGUAUGAGAGAAUGGUAAUUGCCAGUGGCGAAAU ......................................((((((.(((((.....((......))..))))).))))))...(((((...)))))... (-16.93 = -16.93 + 0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:21:10 2011