
| Sequence ID | dm3.chrX |
|---|---|
| Location | 6,399,499 – 6,399,653 |
| Length | 154 |
| Max. P | 0.560059 |

| Location | 6,399,499 – 6,399,653 |
|---|---|
| Length | 154 |
| Sequences | 4 |
| Columns | 173 |
| Reading direction | reverse |
| Mean pairwise identity | 74.85 |
| Shannon entropy | 0.40903 |
| G+C content | 0.46967 |
| Mean single sequence MFE | -44.75 |
| Consensus MFE | -36.39 |
| Energy contribution | -36.45 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.06 |
| Mean z-score | -0.71 |
| Structure conservation index | 0.81 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.560059 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 6399499 154 - 22422827 GGUUAAGACU-CAACCACAGGUGAUUGGUGAAGUGCUCCUUUCGCAUUCGCCAUGGCUGGGAAAAAGCCAUCGAAGAGUGUCAACAGGGGAAAAUCAGUCAGUGCCUGCUUUUCCACCCUUUCC--ACUUGGCAA-------AAAA-----AAAAAACAAAAAAGGACU---- .(((((((((-(.......(((((...((((((......))))))..)))))((((((.......))))))....)))).......((((((((.(((.......))).)))))).))......--.))))))..-------....-----..................---- ( -41.10, z-score = -0.18, R) >droEre2.scaffold_4690 15339892 139 - 18748788 GGUUAAGACU-CCACCACAGGUGAUUGGUGAAGUGCUCCUUUCGCAUUCGCCGUGGCUGGGAUAAAGCCAUCGAAGAGUGUCAACAGGAGAAAAUCAGUCAGUGCCUGGUGCCUUCCACCCUCA--AUUUGAAUA-------AGGGACU------------------------ ((..(((...-.(((((..(.((((((((......(((((((.((((((..(((((((.......))))).))..)))))).)).)))))...)))))))).)...))))).))))).((((..--.........-------))))...------------------------ ( -43.60, z-score = -0.72, R) >droYak2.chrX 2371430 173 - 21770863 GGUUAAGACUGCAACCACAGGUGAUUGGUGAAGUGCUCCUUUCGCAUUCGCCGGGGCUGGAAAAAAGCCAUCAAAGAGUGUCAACAGGAGAAAAUCAGUCAGUGCCUGGUUUUCCUAGAUCUCCUAGUUUUCCUGGUUUUCCACACCCCCCACCCAACUUGAAAAAAGUAACU ((((........))))...((((...((((((.(((.......)))))))))((((.(((((((..(.(((......))).)..((((((((((((((.......)))))))))((((.....))))...))))).)))))))..)))).))))..((((.....)))).... ( -52.60, z-score = -1.56, R) >droSec1.super_4 5786412 159 + 6179234 GGUUAAGACU-CAACCACAGGUGAUUGGUGAAGUGCUCCUUUCGCAUUCGCCGUGGCUGGGAAAAAGCCAUCGAAGAGUGUCAACAGGGGAAAAUCAGUCAGUGCCUGCUUUUCCACCCUUUCC--ACUUCACAA-------AAAAAAAAGAAAAAAAGAAAAAGGACU---- ((((....((-((.((((.(((((...((((((......))))))..))))))))).))))....))))...((((.((....)).((((((((.(((.......))).)))))).))......--.))))....-------...........................---- ( -41.70, z-score = -0.37, R) >consensus GGUUAAGACU_CAACCACAGGUGAUUGGUGAAGUGCUCCUUUCGCAUUCGCCGUGGCUGGGAAAAAGCCAUCGAAGAGUGUCAACAGGAGAAAAUCAGUCAGUGCCUGCUUUUCCACCCUCUCC__ACUUGACAA_______AAAAAC___AAAAAAC_AAAAAGGACU____ ((((........)))).((((((((((((......(((((((.((((((...((((((.......))))))....)))))).)).)))))...)))))))...)))))................................................................. (-36.39 = -36.45 + 0.06)

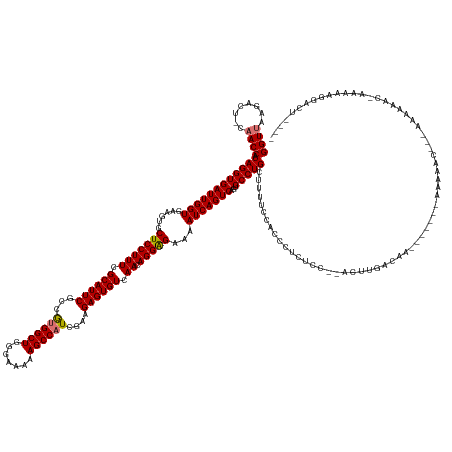

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:20:51 2011