
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 10,440,828 – 10,440,953 |
| Length | 125 |
| Max. P | 0.618455 |

| Location | 10,440,828 – 10,440,953 |
|---|---|
| Length | 125 |
| Sequences | 5 |
| Columns | 136 |
| Reading direction | forward |
| Mean pairwise identity | 82.94 |
| Shannon entropy | 0.27931 |
| G+C content | 0.38119 |
| Mean single sequence MFE | -25.76 |
| Consensus MFE | -19.62 |
| Energy contribution | -19.82 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.76 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.618455 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
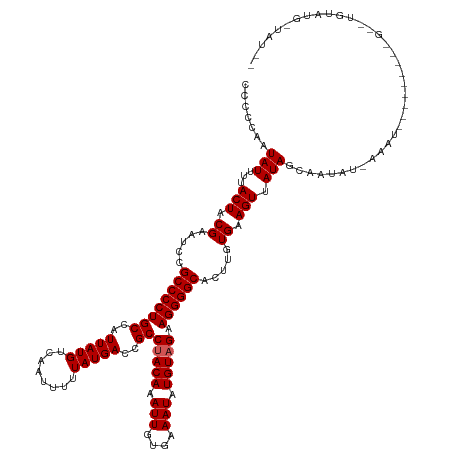
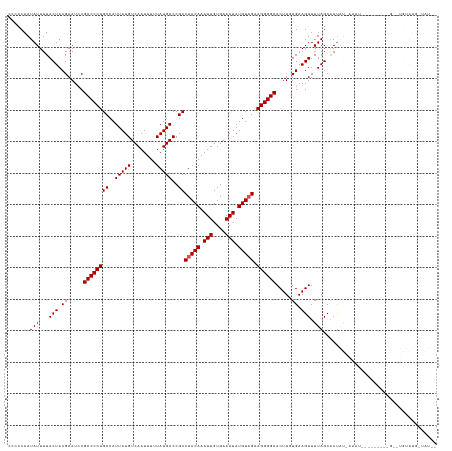
>dm3.chr2L 10440828 125 + 23011544 CCCCCAAUAUUUUACUACGAAUCCGCCCCUGCCAUUAUGUCAAUUUUUAUGACCGCCUACAAAUUGUGAAAUAUGUAGAAGGGGCACUUGUGAAGUUAUAGCAAUAUAAAAU---------GUAUGUAUGUUAU-- ........(((((((...(....)((((((((..(((((........)))))..))(((((.(((....))).))))).))))))....))))))).(((((((((((....---------.))))).))))))-- ( -26.20, z-score = -1.47, R) >droSim1.chr2L 10234967 134 + 22036055 CCCCCAAUAUUUUACUACGAAUCCGCCCCUGCCAUUAUGUCAAUUUUUAUGACCGCCUACAAAUUGUGAAAUAUGUUGAAGGGGCACUUGUGAAGUUAUAGCAAUAUAAAAUAGUUAACUCGGUUGUAUGGUAU-- ......((((..(((..(((....((((((..((.(((((((((((.((........)).))))))....))))).)).))))))..((((....(((((....))))).)))).....)))...)))..))))-- ( -25.80, z-score = 0.22, R) >droSec1.super_3 5868718 136 + 7220098 CCCCCAAUAUUUUACUACGAAUCCGCCCCUGCCAUUAUGUCAAUUUUUAUGACCGCCUACAAAUUGUGAAAUAUGUAGAAGGGGCACUUAUGAAGUUAUAGCGACAUGAAAUAGUUAACUCGCUUGUAUGCUAUAU ........................((((((((..(((((........)))))..))(((((.(((....))).))))).))))))...........((((((((((.(....((....))..).))).))))))). ( -28.00, z-score = -1.39, R) >droYak2.chr2L 6845862 105 + 22324452 CCCCCAAUAUUUUACUACGAAUCCGCCCCUGCCAUUAUGUCAAUUUUUAUGACCGCCUACAAAUUGUGAAAUAUGUAGUAGGGGCACUUUUGAAGUUAUAGCAAU------------------------------- .......(((...(((.((((...((((((((......((((.......))))....((((.(((....))).))))))))))))...)))).))).))).....------------------------------- ( -24.40, z-score = -2.51, R) >droEre2.scaffold_4929 11649780 105 - 26641161 CCCCCAAUAUUUUACUACGAAUCCGCCCCUGCCAUUAUGUCAAUUUUUAUGACCGCCUACAGAUUGUGAAAUAUGUAGUAGGGGCACUUUUGAAGUUAUAGCAAU------------------------------- .......(((...(((.((((...((((((((......((((.......))))....((((.(((....))).))))))))))))...)))).))).))).....------------------------------- ( -24.40, z-score = -2.30, R) >consensus CCCCCAAUAUUUUACUACGAAUCCGCCCCUGCCAUUAUGUCAAUUUUUAUGACCGCCUACAAAUUGUGAAAUAUGUAGAAGGGGCACUUGUGAAGUUAUAGCAAUAU_AAAU_________G__UGUAUG_UAU__ .......(((...(((.((.....((((((((..(((((........)))))..))(((((.(((....))).))))).)))))).....)).))).))).................................... (-19.62 = -19.82 + 0.20)

| Location | 10,440,828 – 10,440,953 |
|---|---|
| Length | 125 |
| Sequences | 5 |
| Columns | 136 |
| Reading direction | reverse |
| Mean pairwise identity | 82.94 |
| Shannon entropy | 0.27931 |
| G+C content | 0.38119 |
| Mean single sequence MFE | -26.72 |
| Consensus MFE | -24.18 |
| Energy contribution | -24.58 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -0.89 |
| Structure conservation index | 0.90 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.596021 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
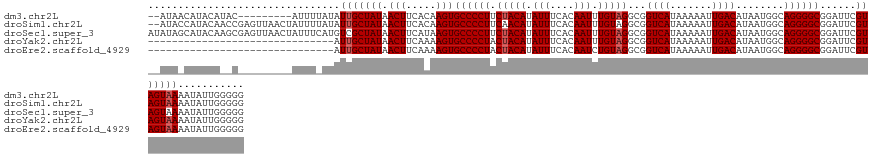
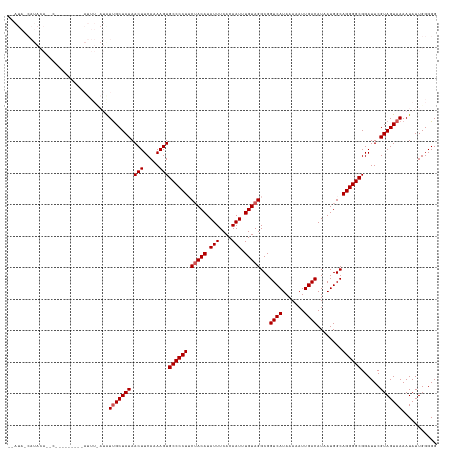
>dm3.chr2L 10440828 125 - 23011544 --AUAACAUACAUAC---------AUUUUAUAUUGCUAUAACUUCACAAGUGCCCCUUCUACAUAUUUCACAAUUUGUAGGCGGUCAUAAAAAUUGACAUAAUGGCAGGGGCGGAUUCGUAGUAAAAUAUUGGGGG --............(---------(...(((.(((((((.(((.....)))((((((((((((.(((....))).))))))..((((.......))))........))))))......))))))).))).)).... ( -26.90, z-score = -1.38, R) >droSim1.chr2L 10234967 134 - 22036055 --AUACCAUACAACCGAGUUAACUAUUUUAUAUUGCUAUAACUUCACAAGUGCCCCUUCAACAUAUUUCACAAUUUGUAGGCGGUCAUAAAAAUUGACAUAAUGGCAGGGGCGGAUUCGUAGUAAAAUAUUGGGGG --...((...(((...((....))....(((.(((((((.(((.....)))((((((((((((.(((....))).))).....((((.......))))....))).))))))......))))))).)))))).)). ( -25.50, z-score = 0.34, R) >droSec1.super_3 5868718 136 - 7220098 AUAUAGCAUACAAGCGAGUUAACUAUUUCAUGUCGCUAUAACUUCAUAAGUGCCCCUUCUACAUAUUUCACAAUUUGUAGGCGGUCAUAAAAAUUGACAUAAUGGCAGGGGCGGAUUCGUAGUAAAAUAUUGGGGG .((((((..(((.(.((((.....))))).))).)))))).(((((....(((((((((((((.(((....))).))))))..((((.......))))........))))))).....(((......)))))))). ( -28.80, z-score = -0.34, R) >droYak2.chr2L 6845862 105 - 22324452 -------------------------------AUUGCUAUAACUUCAAAAGUGCCCCUACUACAUAUUUCACAAUUUGUAGGCGGUCAUAAAAAUUGACAUAAUGGCAGGGGCGGAUUCGUAGUAAAAUAUUGGGGG -------------------------------.(((((((.(((.....)))((((((.(((((.(((....))).)))))...((((.......))))........))))))......)))))))........... ( -26.30, z-score = -1.64, R) >droEre2.scaffold_4929 11649780 105 + 26641161 -------------------------------AUUGCUAUAACUUCAAAAGUGCCCCUACUACAUAUUUCACAAUCUGUAGGCGGUCAUAAAAAUUGACAUAAUGGCAGGGGCGGAUUCGUAGUAAAAUAUUGGGGG -------------------------------.(((((((.(((.....)))((((((.(((((.(((....))).)))))...((((.......))))........))))))......)))))))........... ( -26.10, z-score = -1.44, R) >consensus __AUA_CAUACA__C_________AUUU_AUAUUGCUAUAACUUCAAAAGUGCCCCUUCUACAUAUUUCACAAUUUGUAGGCGGUCAUAAAAAUUGACAUAAUGGCAGGGGCGGAUUCGUAGUAAAAUAUUGGGGG ................................(((((((.(((.....)))((((((.(((((.(((....))).)))))...((((.......))))........))))))......)))))))........... (-24.18 = -24.58 + 0.40)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:30:07 2011