| Sequence ID | dm3.chrX |
|---|---|
| Location | 6,375,103 – 6,375,238 |
| Length | 135 |
| Max. P | 0.928233 |

| Location | 6,375,103 – 6,375,238 |
|---|---|
| Length | 135 |
| Sequences | 6 |
| Columns | 149 |
| Reading direction | forward |
| Mean pairwise identity | 72.39 |
| Shannon entropy | 0.49810 |
| G+C content | 0.41214 |
| Mean single sequence MFE | -32.24 |
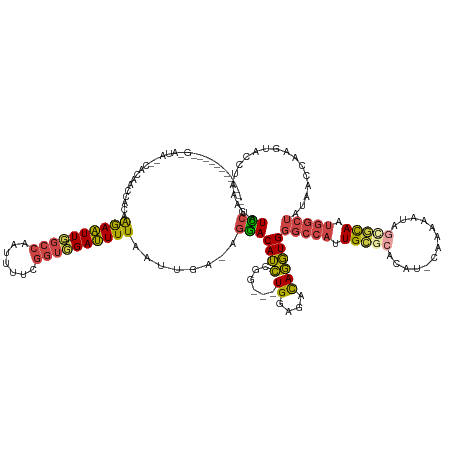
| Consensus MFE | -17.62 |
| Energy contribution | -17.90 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.52 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.34 |
| SVM RNA-class probability | 0.928233 |
| Prediction | RNA |
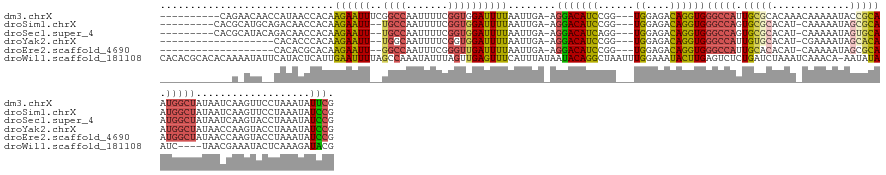
Download alignment: ClustalW | MAF
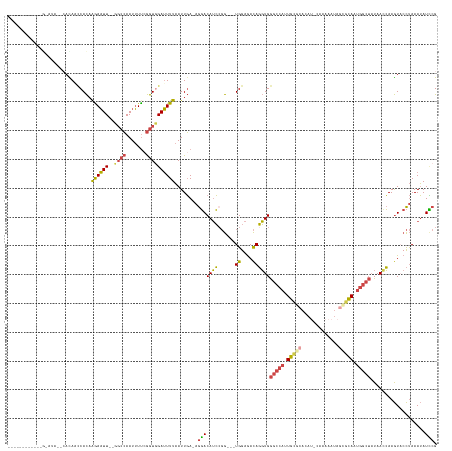
>dm3.chrX 6375103 135 + 22422827 ----------CAGAACAACCAUAACCACAAGAAUUUCGGCCAAUUUUCGGUGGAUUUUAAUUGA-AGGACAUCCGG---UGGAGACAGGUGGGCCAUUGCGCACAAACAAAAAUACCGCAAUGGCUAUAAUCAAGUUCCUAAAUAUUCG ----------..((((........((((.....((((.(((....((((((........)))))-)((....))))---).))))...))))(((((((((...............))))))))).........))))........... ( -37.06, z-score = -2.68, R) >droSim1.chrX 4990749 133 + 17042790 ---------CACGCAUGCAGACAACCACAAGAAUU--UGCCAAUUUUCGGUGGAUUUUAAUUGA-AGGACAUCCGG---UGGAGACAGGUGGGCCAGUGCGCACAU-CAAAAAUAGCGCAAUGGCUAUAAUCAAGUUCCUAAAUAUCCG ---------..............(((...((((((--..((.......))..))))))......-.((....))))---)((((...(((.(((((.(((((....-........))))).)))))...)))...)))).......... ( -35.80, z-score = -1.39, R) >droSec1.super_4 5762725 133 - 6179234 ---------CACGCAUACAGACAACCACAAGAAUU--UGCCAAUUUUCGGUGGAUUUUAAUUGA-AGGACAUCAGG---UGGAGACAGGUGGGCCAGUGCGCACAU-CAAAAAUAGUGCAAUGGCUAUAAUCAAGUACCUAAAUAUCCG ---------....................((((((--..((.......))..))))))......-.(((....(((---((....).(((.(((((.(((((....-........))))).)))))...)))....)))).....))). ( -33.00, z-score = -1.38, R) >droYak2.chrX 2346856 123 + 21770863 -------------------CACACCCACAAGAAUU--UGGCAAUUUUCGGUGGAUUUUAAUUGA-AGGACAUCCGG---UGGAGACAGGUGGGCCAUUGUGCACAU-CGAAAAUAGCACAAUGGCUAUAACCAAGUACCUAAAUAUCCG -------------------.....((((.((((((--....))))))..))))...........-.(((.((..((---((....).(((.(((((((((((....-........)))))))))))...)))....)))...)).))). ( -35.70, z-score = -2.29, R) >droEre2.scaffold_4690 15315634 123 + 18748788 -------------------CACACGCACAAGAAUU--GGCCAAUUUCGGGUUGAUUUUAAUUGA-AGGACAUCCGG---UGGAGACAGGUGGGCCAUUGCACACAU-CAAAAAUAGCGCAAUGGCUAUAACCAAGUACCUAAAUAUCCG -------------------..........(((((.--.(((.......)))..)))))......-.(((.((..((---((....).(((.(((((((((.(....-........).)))))))))...)))....)))...)).))). ( -34.50, z-score = -2.25, R) >droWil1.scaffold_181108 4199134 144 + 4707319 CACACGCACACAAAAUAUUCAUACUCAUUGAAUUUUAGCCAAAUAUUUAGUUGAGUUUCAUUUAUAAUACAGGCUAAUUUGGAAAUACUUGAGUCUCUGAUCUAAAUCAAACA-AAUAUAAUC----UAACGAAAUACUCAAAGAUACG ..........((((..(((((.......))))).((((((...(((((((.((.....)).))).))))..)))))))))).......((((((...((((....))))....-.......((----....))...))))))....... ( -17.40, z-score = -0.42, R) >consensus ____________G_AUA__CACAACCACAAGAAUU__GGCCAAUUUUCGGUGGAUUUUAAUUGA_AGGACAUCCGG___UGGAGACAGGUGGGCCAUUGCGCACAU_CAAAAAUAGCGCAAUGGCUAUAACCAAGUACCUAAAUAUCCG ........................((((..(((...........)))..)))).............(((..........((....))(((.(((((.(((((.............))))).)))))...))).............))). (-17.62 = -17.90 + 0.28)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:20:43 2011