
| Sequence ID | dm3.chrX |
|---|---|
| Location | 6,357,753 – 6,357,826 |
| Length | 73 |
| Max. P | 0.873335 |

| Location | 6,357,753 – 6,357,826 |
|---|---|
| Length | 73 |
| Sequences | 8 |
| Columns | 76 |
| Reading direction | reverse |
| Mean pairwise identity | 80.17 |
| Shannon entropy | 0.39973 |
| G+C content | 0.28497 |
| Mean single sequence MFE | -14.91 |
| Consensus MFE | -7.47 |
| Energy contribution | -7.41 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.45 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.873335 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
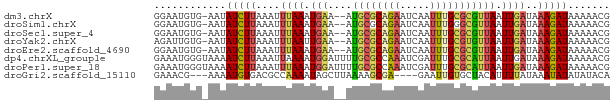
>dm3.chrX 6357753 73 - 22422827 GGAAUGUG-AAUAUCUUAAAUUUAAAUGAA--AUGCGCAGAAUCAAUUUGCGCGUUAAUUGAUAAAGAUAAAAACG .....((.-..((((((....((((....(--(((((((((.....))))))))))..))))..))))))...)). ( -16.10, z-score = -2.50, R) >droSim1.chrX 5001925 73 - 17042790 GGAAUGUG-AAUAUCUUAAAUUUAAAUGAA--AUGCGCAGAAUCAAUUUGGGCGUUAAUUGAUAAAGAUAAAAACG .....((.-..((((((....((((....(--((((.((((.....)))).)))))..))))..))))))...)). ( -12.00, z-score = -1.62, R) >droSec1.super_4 5747949 73 + 6179234 GGAAUGUG-AAUAUCUUAAAUUUAAAUGAA--AUGCGCAGAAUCAAUUUGCGCGUUAAUUGAUAAAGAUAAAAACG .....((.-..((((((....((((....(--(((((((((.....))))))))))..))))..))))))...)). ( -16.10, z-score = -2.50, R) >droYak2.chrX 2333837 73 - 21770863 AGAUUGUG-AAUAUCUUAAAUUUAAUUGAA--AUGCGCAGAAUCAAUUUGCGUGUUAAUUGAUAAAGAUAAAAACG .....((.-..((((((....((((((((.--..(((((((.....))))))).))))))))..))))))...)). ( -17.20, z-score = -3.18, R) >droEre2.scaffold_4690 15302311 73 - 18748788 GGAAUGUG-AAUAUCUUAAAUUUAAAUGAA--AUGCGCAGAAUCAAUUUGCGCGUUAAUUGAUAAAGAUAAAAACG .....((.-..((((((....((((....(--(((((((((.....))))))))))..))))..))))))...)). ( -16.10, z-score = -2.50, R) >dp4.chrXL_group1e 2884466 76 - 12523060 GAAAUGGGUAAAAUCUUAAAUUAAAAUGGAUUUUGCGCCAAAUCGAUUUGCGCAUUAAUUGAUAAAGAUAAAAACG ....((((((((((((...........))))))))).))).(((((((........)))))))............. ( -15.40, z-score = -2.13, R) >droPer1.super_18 1776178 76 - 1952607 GAAAUGGGUAAAAUCUUAAAUUUAAAUGGAUUUUGCGCCAAAUCGAUUUGCGCAUUAAUUGAUAAAGAUAAAAACG ....((((((((((((...........))))))))).))).(((((((........)))))))............. ( -15.40, z-score = -1.99, R) >droGri2.scaffold_15110 7315088 69 - 24565398 GAAACG---AAAAUGUGACGCCAAAAUAGCUUAAAAGCGA----GAAUUGUGCUACAUUUUAUAAAUAUAUAUACA ......---((((((((.(((.......(((....)))..----.....))).))))))))............... ( -10.94, z-score = -1.71, R) >consensus GGAAUGUG_AAUAUCUUAAAUUUAAAUGAA__AUGCGCAGAAUCAAUUUGCGCGUUAAUUGAUAAAGAUAAAAACG ............(((((....((((.(((....((((((((.....))))))))))).))))..)))))....... ( -7.47 = -7.41 + -0.06)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:20:38 2011