
| Sequence ID | dm3.chrX |
|---|---|
| Location | 6,246,550 – 6,246,604 |
| Length | 54 |
| Max. P | 0.996691 |

| Location | 6,246,550 – 6,246,604 |
|---|---|
| Length | 54 |
| Sequences | 6 |
| Columns | 62 |
| Reading direction | forward |
| Mean pairwise identity | 72.29 |
| Shannon entropy | 0.50034 |
| G+C content | 0.44316 |
| Mean single sequence MFE | -12.52 |
| Consensus MFE | -8.57 |
| Energy contribution | -9.93 |
| Covariance contribution | 1.36 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.97 |
| SVM RNA-class probability | 0.996691 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
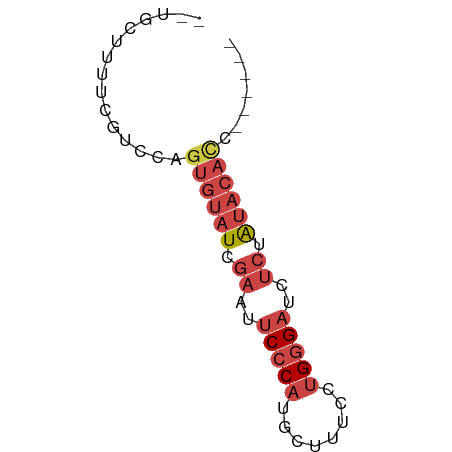
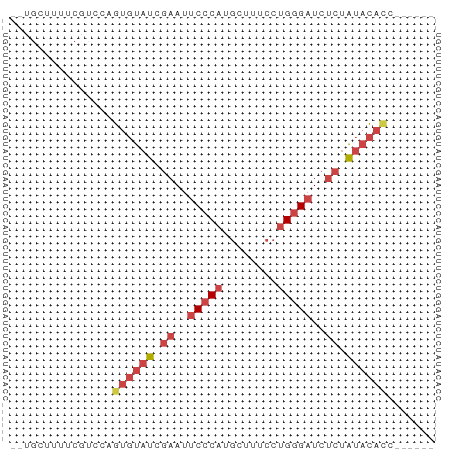
>dm3.chrX 6246550 54 + 22422827 --UGCUUUUCGACCAGUGUAUCGAAUUCCCAUGCUUUCCUGGGAUCUCUAUCCACC------ --.............(((.((.((..(((((........)))))..)).)).))).------ ( -10.90, z-score = -1.47, R) >droSim1.chrX 4919752 54 + 17042790 --UGCUUUUCGUUCAGUGUAUCGAAUUCCCAUGCUUUCCUGGGAUCUCUAUACACC------ --.............((((((.((..(((((........)))))..)).)))))).------ ( -15.50, z-score = -3.97, R) >droSec1.super_4 5655668 54 - 6179234 --UGCUUUUCGUUCAGUGUAUCGAAUUCCCAUGCUUUCCUGGGAUCUCUAUACACC------ --.............((((((.((..(((((........)))))..)).)))))).------ ( -15.50, z-score = -3.97, R) >droYak2.chrX 2240357 58 + 21770863 --UGAUUUUCGUCGAGUGUAUUGAAUUCCCAUGCUUUCCUGGGAUCUCUAUACAUAUAUC-- --.............((((((.((..(((((........)))))..)).)))))).....-- ( -14.20, z-score = -2.31, R) >droEre2.scaffold_4690 15211072 54 + 18748788 --UGCUUUUCGUCCAGUGUAUUGAAUUCCCAUGCUUUCCUGGGAUCUCUAUACAUC------ --.............((((((.((..(((((........)))))..)).)))))).------ ( -13.60, z-score = -3.03, R) >droAna3.scaffold_13335 1205502 60 - 3335858 UGUACUACUCCCCCUAC-UAUUCCACCCACACAGUUUCC-GAGUUUUCCGGAUUGGAUUCCU .................-...((((...........(((-(.......)))).))))..... ( -5.40, z-score = 0.96, R) >consensus __UGCUUUUCGUCCAGUGUAUCGAAUUCCCAUGCUUUCCUGGGAUCUCUAUACACC______ ...............((((((.((..(((((........)))))..)).))))))....... ( -8.57 = -9.93 + 1.36)

| Location | 6,246,550 – 6,246,604 |
|---|---|
| Length | 54 |
| Sequences | 6 |
| Columns | 62 |
| Reading direction | reverse |
| Mean pairwise identity | 72.29 |
| Shannon entropy | 0.50034 |
| G+C content | 0.44316 |
| Mean single sequence MFE | -14.78 |
| Consensus MFE | -7.76 |
| Energy contribution | -9.28 |
| Covariance contribution | 1.53 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.62 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.71 |
| SVM RNA-class probability | 0.994580 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
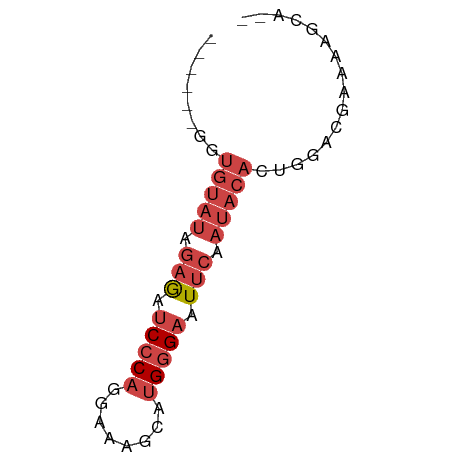
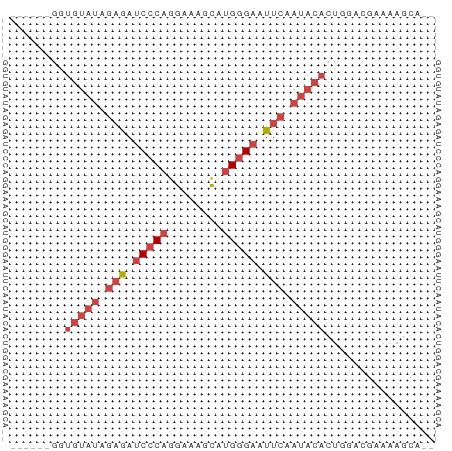
>dm3.chrX 6246550 54 - 22422827 ------GGUGGAUAGAGAUCCCAGGAAAGCAUGGGAAUUCGAUACACUGGUCGAAAAGCA-- ------((((....(((.(((((........))))).)))....))))............-- ( -14.40, z-score = -1.84, R) >droSim1.chrX 4919752 54 - 17042790 ------GGUGUAUAGAGAUCCCAGGAAAGCAUGGGAAUUCGAUACACUGAACGAAAAGCA-- ------(((((((.(((.(((((........))))).))).)))))))............-- ( -18.00, z-score = -4.42, R) >droSec1.super_4 5655668 54 + 6179234 ------GGUGUAUAGAGAUCCCAGGAAAGCAUGGGAAUUCGAUACACUGAACGAAAAGCA-- ------(((((((.(((.(((((........))))).))).)))))))............-- ( -18.00, z-score = -4.42, R) >droYak2.chrX 2240357 58 - 21770863 --GAUAUAUGUAUAGAGAUCCCAGGAAAGCAUGGGAAUUCAAUACACUCGACGAAAAUCA-- --......(((((.(((.(((((........))))).))).)))))..............-- ( -13.80, z-score = -2.66, R) >droEre2.scaffold_4690 15211072 54 - 18748788 ------GAUGUAUAGAGAUCCCAGGAAAGCAUGGGAAUUCAAUACACUGGACGAAAAGCA-- ------..(((((.(((.(((((........))))).))).)))))..............-- ( -13.80, z-score = -2.92, R) >droAna3.scaffold_13335 1205502 60 + 3335858 AGGAAUCCAAUCCGGAAAACUC-GGAAACUGUGUGGGUGGAAUA-GUAGGGGGAGUAGUACA .....(((..(((((.....))-))).(((((.........)))-))....)))........ ( -10.70, z-score = 0.53, R) >consensus ______GGUGUAUAGAGAUCCCAGGAAAGCAUGGGAAUUCAAUACACUGGACGAAAAGCA__ ........(((((.(((.(((((........))))).))).)))))................ ( -7.76 = -9.28 + 1.53)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:20:24 2011