
| Sequence ID | dm3.chrX |
|---|---|
| Location | 6,111,014 – 6,111,145 |
| Length | 131 |
| Max. P | 0.649188 |

| Location | 6,111,014 – 6,111,145 |
|---|---|
| Length | 131 |
| Sequences | 6 |
| Columns | 144 |
| Reading direction | reverse |
| Mean pairwise identity | 66.63 |
| Shannon entropy | 0.63678 |
| G+C content | 0.43018 |
| Mean single sequence MFE | -33.04 |
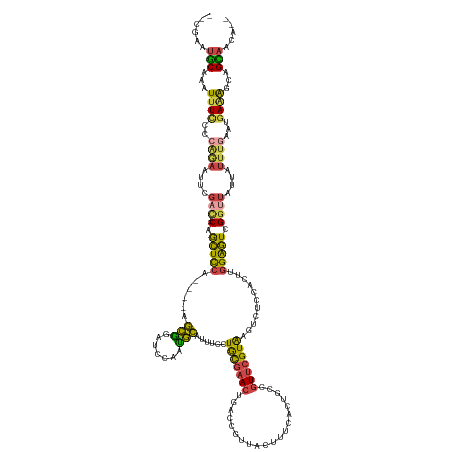
| Consensus MFE | -15.14 |
| Energy contribution | -12.93 |
| Covariance contribution | -2.21 |
| Combinations/Pair | 1.71 |
| Mean z-score | -0.96 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.649188 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
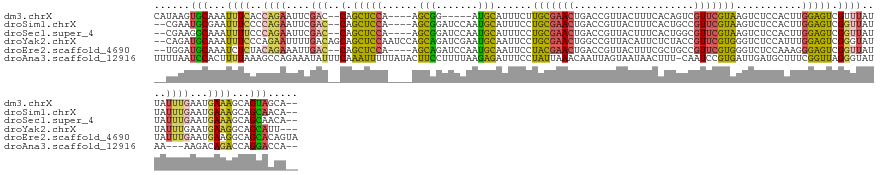
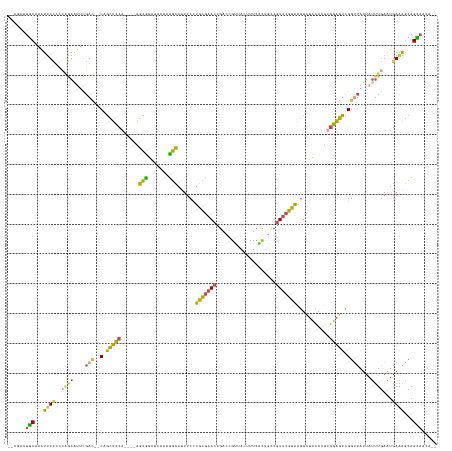
>dm3.chrX 6111014 131 - 22422827 CAUAAGUGCAAAUUUCACCAGAAUUCGAC--CAGCUCCA----AGCGG-----AUGCAUUUCUUGCGAACUGACCGUUACUUUCACAGUCGUUCGUAAGUCUCCACUUGGAGUCGUUUAUUAUUUGAAUGAAAGCAGUAGCA-- ....(.(((...(((((...((((..(((--..((((((----((.((-----(.......(((((((((.(((.((.......)).))))))))))))..))).)))))))).)))....))))...)))))))).)....-- ( -37.00, z-score = -2.80, R) >droSim1.chrX 4826500 134 - 17042790 --CGAAUGCGAAUUUCCCCAGAAUUCGAC--CAGCUCCA----AGCGGAUCCAAUGCAUUUCCUGCGAACUGACCGUUACUUUCACUGCCGUUCGUAAGUCUCCACUUGGAGUCGGUUAUUAUUUGAAUGAAAGCAGCAACA-- --....(((...((((....((((..(((--(.((((((----((.(((..(...(((.....)))((((.(.(.((.......)).)).))))....)..))).)))))))).))))...))))....))))...)))...-- ( -36.70, z-score = -2.15, R) >droSec1.super_4 5520500 134 + 6179234 --CGAAGGCAAAUUUUCCCAGAAUUCGAC--CAGCUCCA----AGCGGAUCCAAUGCAUUUCCUGCGAACUGACCGUUACUUUCACUGGCGUUCGUAAGUCUCCACUUGGAGUCGGUUAUUAUUUGAAUGAAAGCAGCAACA-- --.....((...(((((...((((..(((--(.((((((----((.(((..(...(((.....)))((((.(.(((..........))))))))....)..))).)))))))).))))...))))....)))))..))....-- ( -34.90, z-score = -1.02, R) >droYak2.chrX 2115357 139 - 21770863 --CAGAUGCAAAUUUCCCAGAAUUUUGACAGCAGCUCCAAUCCAGCAGAUCGAAUGCAAUUCCUGCGAACUGGCCGUUACAUUCUCUACCGUUCGUGGGUCUCCAUUUGGAGUCGGGUAUUAUUUGAAUGAAGGCAGCAUU--- --..(((((...((((...((((....((..(.(((((((....(((.......)))....((..(((((.((...............)))))))..)).......))))))).).))...))))....))))...)))))--- ( -32.96, z-score = 0.68, R) >droEre2.scaffold_4690 15081252 136 - 18748788 --UGGAUGCAAAUCUCUACAGAAAUUGAC--CAGCUCCA----AGCAGAUCCAAUGCAAUUCCUACGAACUGACCGUUACUUUCGCUGCCGUUCGUGGGUCUCCAAAGGGAGUCGGUUAUUAUUUGAAUGAAGGCAGCACAGUA --((..(((...((....((((...((((--(.(((((.----.(((.......)))....(((((((((.(..((.......))...).))))))))).........))))).)))))...))))...))..)))...))... ( -37.20, z-score = -0.79, R) >droAna3.scaffold_12916 13493036 138 + 16180835 UUUUAAUCCACUUUUAAAGCCAGAAAUAUUUCAAAUUUUUAUACUUCCUUUUAAGAGAUUUCCUAUUAAACAAUUAGUAAUAACUUU-CAAUCCGUGAUUGAUGCUUUCGGUUAGGGUAUAA---AAGACAGACCAGGACCA-- ......(((.(((((((((...(((.(((...........))).))).)))))))))......((((((....))))))....((((-....((.(((((((.....))))))).))....)---)))........)))...-- ( -19.50, z-score = 0.32, R) >consensus __CGAAUGCAAAUUUCCCCAGAAUUCGAC__CAGCUCCA____AGCGGAUCCAAUGCAUUUCCUGCGAACUGACCGUUACUUUCACUGCCGUUCGUAAGUCUCCACUUGGAGUCGGUUAUUAUUUGAAUGAAAGCAGCAACA__ ......(((...((((............................(((.......)))....(((((((((....................))))))).(.((((....)))).))).............))))...)))..... (-15.14 = -12.93 + -2.21)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:19:56 2011