
| Sequence ID | dm3.chrX |
|---|---|
| Location | 6,008,000 – 6,008,063 |
| Length | 63 |
| Max. P | 0.810399 |

| Location | 6,008,000 – 6,008,063 |
|---|---|
| Length | 63 |
| Sequences | 13 |
| Columns | 63 |
| Reading direction | forward |
| Mean pairwise identity | 79.08 |
| Shannon entropy | 0.48350 |
| G+C content | 0.61294 |
| Mean single sequence MFE | -25.14 |
| Consensus MFE | -16.10 |
| Energy contribution | -16.16 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.65 |
| Mean z-score | -1.19 |
| Structure conservation index | 0.64 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.810399 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 6008000 63 + 22422827 AUCCCGGGCGAGCCGGAUCUUGGAGAGGCGUGGCGCCUCGCUGCGCCGCUCCUUGAUGGAGUA ...((((.....))))(((..((((.((((..(((...)))..)))).))))..)))...... ( -32.20, z-score = -1.92, R) >droSim1.chrX_random 1896919 63 + 5698898 AUCCCGGGCGAGCCGGAUUUUGGAGAGGCGUGGCGCCUCGCUGCGCCGCUCCUUGAUGGAGUA ...((((.....))))(((..((((.((((..(((...)))..)))).))))..)))...... ( -30.40, z-score = -1.71, R) >droSec1.super_4 5421381 63 - 6179234 AUCCCGGGCGAGCCGGAUUUUGGAGAGGCGUGGCGCCUCGCUGCGCCGCUCCUUGAUGGAGUA ...((((.....))))(((..((((.((((..(((...)))..)))).))))..)))...... ( -30.40, z-score = -1.71, R) >droYak2.chrX 2017886 63 + 21770863 AUCCCGGGCGAGCCGGAUCUUGGAGAGGCGUGGCGCCUCGCUGCGCCGCUCCUUGAUGGAGUA ...((((.....))))(((..((((.((((..(((...)))..)))).))))..)))...... ( -32.20, z-score = -1.92, R) >droEre2.scaffold_4690 14987081 63 + 18748788 AUCCCGGGCGAGCCGGAUCUUGGAGAGGCGUGGCGCCUCGCUGCGCCGCUCCUUGAUGGAGUA ...((((.....))))(((..((((.((((..(((...)))..)))).))))..)))...... ( -32.20, z-score = -1.92, R) >droAna3.scaffold_13417 604370 63 + 6960332 AUCCCGGACGAGACGGAUUUUGGAGAGGCGUGGCGCCUCGCUGCGUCGCUCCUUGAUCGAGUA ....((.......))((((..((((.((((..(((...)))..)))).))))..))))..... ( -25.50, z-score = -1.10, R) >dp4.Unknown_group_804 53530 63 + 58332 CUCCCGGACGAGCCGGAUCUUGGAAAGGCGCGACGCCUCGCUGCGUCUGUCCUUGACGGAGUA (((((((.....)))..((..(((.(((((((.((...)).))))))).)))..)).)))).. ( -23.90, z-score = -0.60, R) >droPer1.super_324 22913 63 - 26204 CUCCCGGACGAGCCGGAUCUUGGAAAGGCGCGGCGCCUCGCUGCGUCUGUCCUUGACGGAGUA (((((((.....)))..((..(((.((((((((((...)))))))))).)))..)).)))).. ( -30.50, z-score = -2.25, R) >droWil1.scaffold_181150 2254741 63 - 4952429 UUCGCGUACCAACCGGAUUUUGGAGAGGCGCGAAGCAUCGCCCCGCCUGUCCUUGAUGGAGUA ............(((......(((.((((((((....))))...)))).)))....))).... ( -18.50, z-score = -0.18, R) >droVir3.scaffold_12928 730959 63 + 7717345 AUCGCGUAUGAGCCGCAUUUUGGUUAGGCGUGGCGCCUCGCUGCGUCGUUCCUUGAUCGAGUA ...((......))((.(((..((...((((..(((...)))..))))...))..))))).... ( -19.80, z-score = -0.01, R) >droMoj3.scaffold_6328 1700078 63 + 4453435 AUCGCGUAUGAGACGCAUUUUCGAUAACCGUGGCGCCUCGUUGCGUCGCUCCUUGAUCGAGUA ...((((.....))))...((((((....(((((((......)))))))......)))))).. ( -20.20, z-score = -1.02, R) >droGri2.scaffold_14853 2382182 63 - 10151454 AUCGCGUAUAAGUCGCAUCUUGGAUAUGCGUGGCGCCUCGUUGCGUCGAUCCUUGAUCGAGUA ..........(.(((.(((..((((..(((..(((...)))..)))..))))..)))))).). ( -20.60, z-score = -0.99, R) >anoGam1.chr2R 22074779 56 + 62725911 AUCCUGCAC---CUGGAUGUUGUACAGCUUUAAAGAUUC----CGCCUUUUUCUGGGUGAAUA ......(((---(..((.(((....)))...((((....----...)))).))..)))).... ( -10.40, z-score = -0.16, R) >consensus AUCCCGGACGAGCCGGAUCUUGGAGAGGCGUGGCGCCUCGCUGCGCCGCUCCUUGAUGGAGUA ...(((.......))).((..(((..(((((((((...)))))))))..)))..))....... (-16.10 = -16.16 + 0.06)


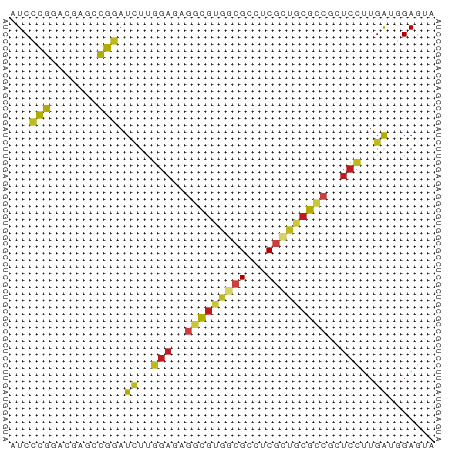
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:19:44 2011