
| Sequence ID | dm3.chrX |
|---|---|
| Location | 5,931,541 – 5,931,591 |
| Length | 50 |
| Max. P | 0.996580 |

| Location | 5,931,541 – 5,931,591 |
|---|---|
| Length | 50 |
| Sequences | 5 |
| Columns | 55 |
| Reading direction | reverse |
| Mean pairwise identity | 60.96 |
| Shannon entropy | 0.67204 |
| G+C content | 0.44000 |
| Mean single sequence MFE | -11.60 |
| Consensus MFE | -4.48 |
| Energy contribution | -4.12 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.67 |
| Mean z-score | -2.56 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.95 |
| SVM RNA-class probability | 0.996580 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
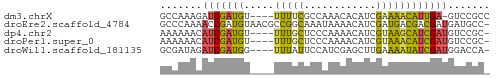
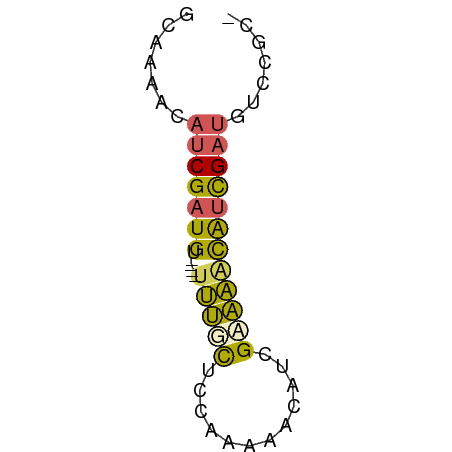
>dm3.chrX 5931541 50 - 22422827 GCCAAAGAUCGAUGU----UUUUCGCCAAACACAUCGAAAACAUUGA-GUCCGCC ((......(((((((----(((.((..........))))))))))))-....)). ( -9.30, z-score = -1.66, R) >droEre2.scaffold_4784 12344272 54 + 25762168 GCCCAAAACCGAUGUAACGCCGGCAAAUAAAACAUCGAUGACGACGAUGAUGCC- (((......((......))..)))........(((((.......))))).....- ( -7.00, z-score = -0.20, R) >dp4.chr2 8371470 50 - 30794189 AAAAAACAUCGAUGU----UUUGCUCCCAAAACAUCGUAAGCAUCGAUGUCCGC- .....((((((((((----((.((............))))))))))))))....- ( -14.10, z-score = -4.40, R) >droPer1.super_0 2206045 50 - 11822988 AAAAAACAUCGAUGU----UUUGCUCCCAAAACAUCGUAAACAUCGAUGUCCGC- .....((((((((((----((.((............))))))))))))))....- ( -14.10, z-score = -4.87, R) >droWil1.scaffold_181135 13205 50 - 74897 GCGAUAGAUCGAUGG----UUUAUUCCAUCGAGCUUGAAAAUAUCGAUGGACCA- .(((((..(((((((----......))))))).........)))))........- ( -13.50, z-score = -1.70, R) >consensus GCAAAACAUCGAUGU____UUUGCUCCAAAAACAUCGAAAACAUCGAUGUCCGC_ .......((((((((.....((((............))))))))))))....... ( -4.48 = -4.12 + -0.36)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:19:26 2011