
| Sequence ID | dm3.chrX |
|---|---|
| Location | 5,723,468 – 5,723,572 |
| Length | 104 |
| Max. P | 0.837169 |

| Location | 5,723,468 – 5,723,572 |
|---|---|
| Length | 104 |
| Sequences | 7 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 74.44 |
| Shannon entropy | 0.43682 |
| G+C content | 0.52643 |
| Mean single sequence MFE | -34.14 |
| Consensus MFE | -19.01 |
| Energy contribution | -19.00 |
| Covariance contribution | -0.01 |
| Combinations/Pair | 1.28 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.86 |
| SVM RNA-class probability | 0.837169 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
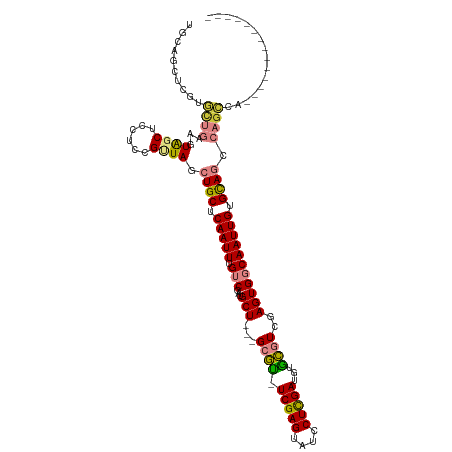
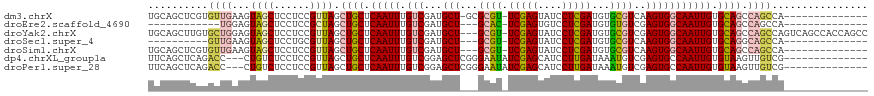
>dm3.chrX 5723468 104 - 22422827 UGCAGCUCGUGUUGAAGUAGCUCCUCCGUUAGCUGCUCAAUUUGUCGAUGCU-GCGCGU-UCGAGUAUCCUCGAUGUGCGUCAAGUGGCAAUUGUGCAGCCAGCCA-------------- ..((((....)))).............(((.(((((.(((((.((((.((.(-((((((-.((((....))))))))))).))..))))))))).))))).)))..-------------- ( -36.30, z-score = -1.65, R) >droEre2.scaffold_4690 3071535 90 - 18748788 ------------UGGAGUAGCUCCUCCGCUAGCUGCUCAAUUUGUCGAUGCU---GCAC-UCGAGUGUCCUCGAUGUGUGUCGAGUGGCAAUUGUGCAGCCAGCCA-------------- ------------.((((......))))(((.(((((.(((((.((((.((.(---((((-(((((....))))).))))).))..))))))))).))))).)))..-------------- ( -37.30, z-score = -3.03, R) >droYak2.chrX 19021874 116 + 21770863 UGCAGCUUGUGCUGGAGUAGCUCCUCCGUUAGCUGCUCAAUUUGUCGAUGCU---GCGU-UCGAGUAUCCUCGAUGUGCGUCGAGUGGCAAUUGUGCAGCCAGCCAGUCAGCCACCAGCC ..((((....))))((((((((........))))))))......(((((((.---((((-.((((....)))))))))))))))(((((.((((.((.....))))))..)))))..... ( -46.90, z-score = -2.88, R) >droSec1.super_4 5137189 92 + 6179234 ----------GUUGAAGUAGCUCCUGCGUUAGCUGCUCAAUUUGUCGAUGCU---GCGU-UCGAGUAUCCUCGAUGUGCGUCAAGUGGCAAUUGUGCAGGCAGCCA-------------- ----------(((((.((((((........)))))))))))((((((((((.---((((-.((((....))))))))))))).....(((....))).)))))...-------------- ( -30.70, z-score = -0.99, R) >droSim1.chrX 4461971 102 - 17042790 UGCAGCUCGUGUUGAAGUAGCUCCUCCGUUAGCUGCUCAAUUUGUCGAUGCU---GCGU-UCGAGUAUCCUCGAUGUGCGUCAAGUGGCAAUUGUGCAGCCAGCCA-------------- ..((((....)))).............(((.(((((.(((((.((((((((.---((((-.((((....)))))))))))))....)))))))).))))).)))..-------------- ( -33.40, z-score = -1.34, R) >dp4.chrXL_group1a 7069236 103 + 9151740 UUCAGCUCAGACC---CUGUCUCCUCCGUUAGCUGCUCAAUUUGUCGGAGCUCGGGAAUAUCGAGCAUCCUUGAUAAAUGUCGAGUGCCAAUUGUGUAAGUUGUCG-------------- ..(((((.((((.---..)))).(.(.(((.((.((((.((((((((((((((((.....)))))).)))..)))))))...)))))).))).).)..)))))...-------------- ( -27.20, z-score = -1.23, R) >droPer1.super_28 595387 103 + 1111753 UUCAGCUCAGACC---CUGUCUCCUCCGUUAGCUGCUCAAUUUGUCGGAGCUCGGGAAUAUCGAGCAUCCUUGAUAAAUGUCGAGUGCCAAUUGUGUAAGUUGUCG-------------- ..(((((.((((.---..)))).(.(.(((.((.((((.((((((((((((((((.....)))))).)))..)))))))...)))))).))).).)..)))))...-------------- ( -27.20, z-score = -1.23, R) >consensus UGCAGCUCGUGCUGAAGUAGCUCCUCCGUUAGCUGCUCAAUUUGUCGAUGCU___GCGU_UCGAGUAUCCUCGAUGUGCGUCGAGUGGCAAUUGUGCAGCCAGCCA______________ ..........((((...((((......)))).((((.(((((.(((...(((...((((.(((((....)))))...))))..))))))))))).)))).))))................ (-19.01 = -19.00 + -0.01)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:18:45 2011