
| Sequence ID | dm3.chrX |
|---|---|
| Location | 5,664,758 – 5,664,856 |
| Length | 98 |
| Max. P | 0.787308 |

| Location | 5,664,758 – 5,664,856 |
|---|---|
| Length | 98 |
| Sequences | 7 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 73.27 |
| Shannon entropy | 0.51010 |
| G+C content | 0.51114 |
| Mean single sequence MFE | -29.03 |
| Consensus MFE | -16.55 |
| Energy contribution | -17.31 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.35 |
| Mean z-score | -1.26 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.787308 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 5664758 98 - 22422827 -----AGCCGCCUUGCUGCCGGCGUCGCUGUCGUUUCCAUUUCCAAUUUCCAUUUACGCGACGAUCGCUU-AAAAGAUCGGUCGUCGGUCAGC-GCA-AACAUUUA---- -----.((((.(.....).))))(.(((((.(((.....................)).(((((((((.((-....)).)))))))))).))))-)).-........---- ( -27.80, z-score = -1.02, R) >droSim1.chrX 4405811 98 - 17042790 -----UGCCGCCUUGCUGCUGGCGUCGCUGUCGUUUCCAUUUCCAAUUUCCAUUUACGCGACGAUCGCUU-AAAAGAUCGGUCGUCGGUCAGC-GCA-AACAUUUA---- -----(((((((........))))..((((.(((.....................)).(((((((((.((-....)).)))))))))).))))-)))-........---- ( -27.50, z-score = -0.91, R) >droSec1.super_4 5080034 98 + 6179234 -----UGCCGCCUUGCUGCUGGCGUCGCUGUCGUUUCCAUUUCCAAUUUCCAUUAACGCGACGAUCGCUU-AAAAGAUCGGUCGUCGGUCAGC-GCA-AACAUUUA---- -----(((((((........))))..((((.((((...................))).(((((((((.((-....)).)))))))))).))))-)))-........---- ( -28.51, z-score = -1.21, R) >droYak2.chrX 18962011 98 + 21770863 -----UGCCGCCUUGCUGCCGGCGUCGCUGUCGUUUCCAUUUCCAAUUUCCAUUUACGCGACGAUCGCUU-UAAAGAUCGGUCGUCGGUCAGC-GCA-AACAUUUA---- -----.((((.(.....).))))(.(((((.(((.....................)).(((((((((.((-....)).)))))))))).))))-)).-........---- ( -27.70, z-score = -0.92, R) >droEre2.scaffold_4690 3013220 98 - 18748788 -----UGCCGCCUUGCUGCCGGCGUCGCCGUCGUUUCCAUUUCCAAUUUCCAUUUACGCGACGAUCGCUU-UAAAGAUCGGUCGUCGGUCAGC-GCA-AACAUUUA---- -----....((...((((((((((..((((((((.......................)))))((((....-....))))))))))))).))))-)).-........---- ( -30.10, z-score = -1.66, R) >dp4.chr2 26404479 93 + 30794189 --------UGCCUCCAUCUUGGAGGAGU--CUGCCUCCGUUUUU-GCAUCGACUGCCUCGACGUUUUCACGCUUCGAAUGGCAGUGGGAACGC-ACA-AAUAUUUA---- --------............(((((...--...)))))((((.(-((.((.((((((((((.((......)).))))..)))))).))...))-).)-))).....---- ( -28.40, z-score = -1.59, R) >droWil1.scaffold_181096 5050720 104 + 12416693 UGUUUGGAAGCGUUAACGUCAGCGCUGCUGCGACUGUCACUUC----AUUCAGU-ACGCGACGAUCGCUC-CAAAGAUCAGUCGUUGGACAGCAGCGCAAAAUUUAUUUA .........(((....)))..((((((((((.((((.......----...))))-..(((((((((....-....)))).)))))..).)))))))))............ ( -33.20, z-score = -1.49, R) >consensus _____UGCCGCCUUGCUGCCGGCGUCGCUGUCGUUUCCAUUUCCAAUUUCCAUUUACGCGACGAUCGCUU_AAAAGAUCGGUCGUCGGUCAGC_GCA_AACAUUUA____ .........((...((((((((((((((.............................)))))((((.........))))....))))).)))).)).............. (-16.55 = -17.31 + 0.76)

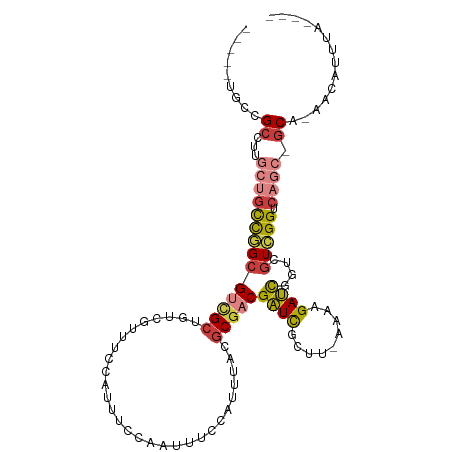

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:18:32 2011