
| Sequence ID | dm3.chrX |
|---|---|
| Location | 5,660,932 – 5,661,057 |
| Length | 125 |
| Max. P | 0.987342 |

| Location | 5,660,932 – 5,661,034 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 68.99 |
| Shannon entropy | 0.53774 |
| G+C content | 0.41320 |
| Mean single sequence MFE | -24.90 |
| Consensus MFE | -13.54 |
| Energy contribution | -13.98 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.28 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.27 |
| SVM RNA-class probability | 0.987342 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
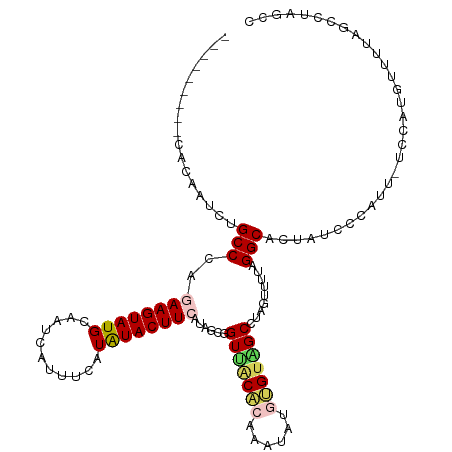
>dm3.chrX 5660932 102 - 22422827 GCCCACUGCCACAAUCUGCCCAGAAGUAUGCUAUCAUUUCAUAUACUUCAUGCUGGUUACACAAAUAUGUGUGGCCUAGUUUUAGGCAUUAUCCUAUU-UCUA--------------- ......((((............((((((((...........))))))))..((((((((((((....))))))))).)))....))))..........-....--------------- ( -26.00, z-score = -2.41, R) >droSim1.chrX 4401512 108 - 17042790 ---------CACAAUCUGCCCAAAAGUAUGCAAUCAUUUCAUAUACUUCAUAGCAGUUGCUCAAAUAUGUGUAGCCUAGUUUUAGGCACUAUCUCAUU-UCCAUGUUUUAGCCUAGCC ---------......((((....(((((((...........)))))))....))))..(((.(((((((.((.(((((....))))))).........-..))))))).)))...... ( -19.20, z-score = -1.16, R) >droSec1.super_4 5076179 108 + 6179234 ---------CACAAUCUGCCCAAAAGUAUGCAAUCAUUUCAUAUACUUCAUAGCAGUUGCACAAAUAUGUGUAGCCUAGUUUUAGGCACUAUCUCAUU-UCCAUGUUUUAGCCUAGCC ---------......((((....(((((((...........)))))))....)))).((((((....))))))(((((....))))).(((...(((.-...)))...)))....... ( -19.40, z-score = -1.11, R) >droYak2.chrX 18958273 105 + 21770863 -----------CGAAAGGCCCAGAAGUAUGCAAUCAUUUCAUAUACUUCUCAGCGGUUACCAAAAUA--GGUUGCCUACUUUUAGGCACCGUGGCAUUGUCCAUCGUCUAGCGUAACU -----------(....)((..(((((((((...........)))))))))..))(((((((......--(((.(((((....))))))))((((......))))......).)))))) ( -28.20, z-score = -2.25, R) >droEre2.scaffold_4690 3009714 105 - 18748788 -----------CCAAAGGCCCAGAAGUAUGCAGUCAUCUCAUGUACUUCACAGCGGUGACCCAAACA--GGCUGCAUACUUUUUGGCACUAUGCCGUUUUUCCAAUCUACGCAUAUAA -----------......(((.((((((((((((((......(((.....)))(.(....).).....--)))))))))))))).)))..(((((.((...........)))))))... ( -31.70, z-score = -3.45, R) >consensus _________CACAAUCUGCCCAGAAGUAUGCAAUCAUUUCAUAUACUUCAUAGCGGUUACACAAAUAUGUGUAGCCUAGUUUUAGGCACUAUCCCAUU_UCCAUGUUUUAGCCUAGCC .................(((..((((((((...........))))))))......(((((((......))))))).........)))............................... (-13.54 = -13.98 + 0.44)


| Location | 5,660,954 – 5,661,057 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 74.22 |
| Shannon entropy | 0.32987 |
| G+C content | 0.36402 |
| Mean single sequence MFE | -21.47 |
| Consensus MFE | -16.01 |
| Energy contribution | -16.35 |
| Covariance contribution | 0.34 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.19 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.509684 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 5660954 103 + 22422827 ACUAGGCCACACAUAUUUGUGUAACCAGCAUGAAGUAUAUGAAAUGAUAGCAUACUUCUGGGCAGAUUGUGGCAGUGGGCAAUUGUUAUGAAGCAAUUAUCUU .....(((((((((....)))).....((..(((((((.............)))))))...)).....)))))...((..(((((((....)))))))..)). ( -24.72, z-score = -0.01, R) >droSim1.chrX 4401549 81 + 17042790 ACUAGGCUACACAUAUUUGAGCAACUGCUAUGAAGUAUAUGAAAUGAUUGCAUACUUUUGGGCAGAUUGUGGCAAUUAUUU---------------------- .....((((((((....)).....(((((..(((((((..((.....))..)))))))..)))))..))))))........---------------------- ( -19.60, z-score = -1.98, R) >droSec1.super_4 5076216 81 - 6179234 ACUAGGCUACACAUAUUUGUGCAACUGCUAUGAAGUAUAUGAAAUGAUUGCAUACUUUUGGGCAGAUUGUGGCAAUUAUCU---------------------- .....((((((((....)))....(((((..(((((((..((.....))..)))))))..)))))...)))))........---------------------- ( -20.10, z-score = -1.58, R) >consensus ACUAGGCUACACAUAUUUGUGCAACUGCUAUGAAGUAUAUGAAAUGAUUGCAUACUUUUGGGCAGAUUGUGGCAAUUAUCU______________________ .....((((((((....)).....(((((..(((((((.............)))))))..)))))..)))))).............................. (-16.01 = -16.35 + 0.34)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:18:31 2011