| Sequence ID | dm3.chrX |
|---|---|
| Location | 5,657,432 – 5,657,540 |
| Length | 108 |
| Max. P | 0.574663 |

| Location | 5,657,432 – 5,657,540 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 77.22 |
| Shannon entropy | 0.42104 |
| G+C content | 0.44281 |
| Mean single sequence MFE | -29.98 |
| Consensus MFE | -18.18 |
| Energy contribution | -19.10 |
| Covariance contribution | 0.92 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.31 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.17 |
| SVM RNA-class probability | 0.574663 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 5657432 108 + 22422827 --GGCAGAAAAAA--AAUAUCAAAAAGAA---AUCUGGGGGCGGAGGAAGUUGGGUGGGGUGGCUGGUUAGGUCCACAGUCGCUGCAUAUUUUCGUGUUUUCUUUCUUUUUCUGU--- --.(((((((((.--..........((..---..))(..((((..((((((...(((.((((((((..........)))))))).))))))))).))))..)....)))))))))--- ( -31.10, z-score = -2.04, R) >droSim1.chrX 4398003 111 + 17042790 --GGCAGAAAAAA-AAAUAUACAAAAAAA-UAAUCUGGGGGCGGAGGAAGUUGGGUGGGGUGGCUGGUUAAGCCCACAGUCGCUGCAUAUUUUCGUGUUUUCUUUCUUUUUCUGU--- --.(((((((((.-(((.((((.......-...((((....))))((((((...(((.((((((((..........)))))))).)))))))))))))....))).)))))))))--- ( -31.00, z-score = -1.73, R) >droSec1.super_4 5072640 112 - 6179234 --GGCAGAAAAAA-AAAUAUACAAAAAAAAUUAUCGGGGGGCGGAGGAAGUUGGGUGGGGUGGCUGGUUAAGCCCACAGUCGCUGCAUAUUUUCGUGUUUUCUUUCUUUUUCUGU--- --.(((((((((.-((...................((..((((..((((((...(((.((((((((..........)))))))).))))))))).))))..)))).)))))))))--- ( -29.70, z-score = -1.27, R) >droYak2.chrX 18954838 110 - 21770863 --GGCAGAAAAAACAAAUAUCAAAAAAAA---AUCUGGGGGCGGAGGAAGUGGGGUGGGGUGGCUAGUUAAGCCCACAGUCGCUGCAUAUUUUCGUGGUUUCUUUUUCUUUCUGU--- --.(((((((...................---....((..((.(((((.(((.(((((.((((((.....)).))))..))))).))).)))))...))..)).....)))))))--- ( -26.59, z-score = -0.13, R) >droEre2.scaffold_4690 3006232 107 + 18748788 --GGCAGAGAAAC-CAAUAUGAAAAAAAA----UCUGUGGGCGGAGGAAGUUGGGUGGGGUGGCUAGUUAAGCCUACAGUCGCUGCAUAUUUUCGUGUUUUCUUUCUUUU-CUGU--- --.((((((((..-.((((((((((....----((((....)))).........(((.(((((((.((.......))))))))).))).)))))))))).......))))-))))--- ( -28.80, z-score = -0.78, R) >droAna3.scaffold_13117 582239 103 - 5790199 UCGGUUGAAAAAA-------ACAGGAAAA------UGGAAAGGGGUGGAGUAAGG-GGGGUAACUAAUUA-GUCCCCACUCGCUGCAUAUUUCCACACAGCUCGACUUUCCGGGCUAU ((.(((......)-------))..))...------((((((...(((.((..((.-((((..(((....)-)))))).))..)).))).))))))...((((((......)))))).. ( -32.70, z-score = -1.89, R) >consensus __GGCAGAAAAAA_AAAUAUAAAAAAAAA___AUCUGGGGGCGGAGGAAGUUGGGUGGGGUGGCUAGUUAAGCCCACAGUCGCUGCAUAUUUUCGUGUUUUCUUUCUUUUUCUGU___ ...(((((((((.....................((((....)))).((((....(((.(((((((............))))))).)))..))))............)))))))))... (-18.18 = -19.10 + 0.92)

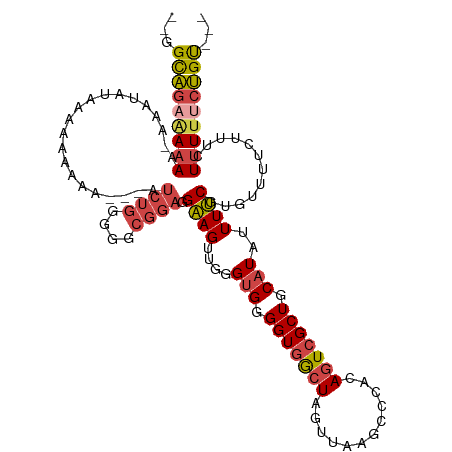
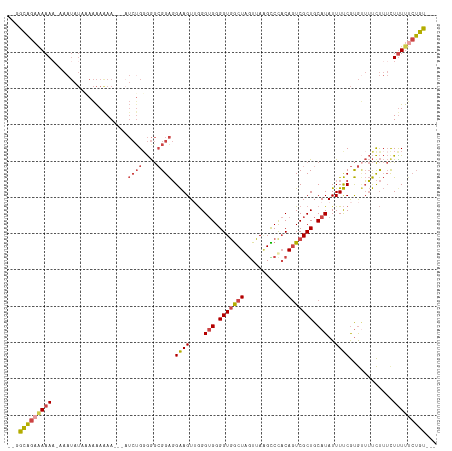
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:18:30 2011