
| Sequence ID | dm3.chrX |
|---|---|
| Location | 5,486,837 – 5,486,932 |
| Length | 95 |
| Max. P | 0.855268 |

| Location | 5,486,837 – 5,486,932 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 74.34 |
| Shannon entropy | 0.47230 |
| G+C content | 0.40994 |
| Mean single sequence MFE | -25.15 |
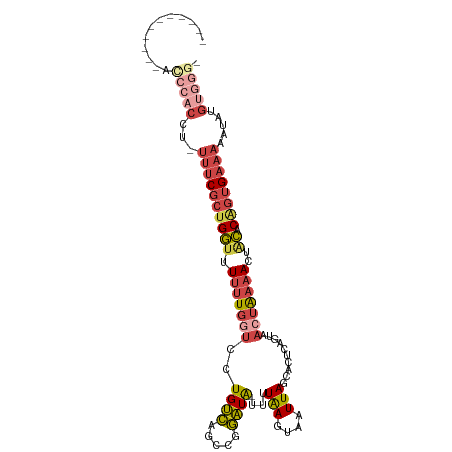
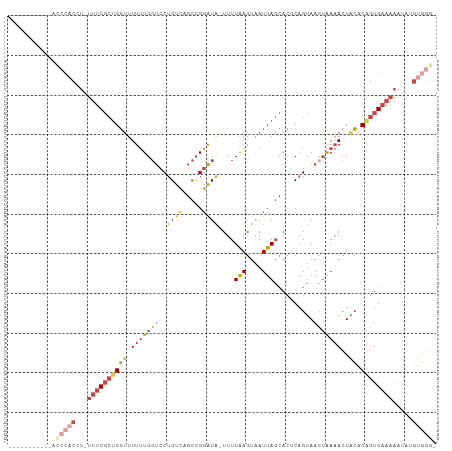
| Consensus MFE | -10.85 |
| Energy contribution | -13.02 |
| Covariance contribution | 2.17 |
| Combinations/Pair | 1.23 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.43 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.93 |
| SVM RNA-class probability | 0.855268 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
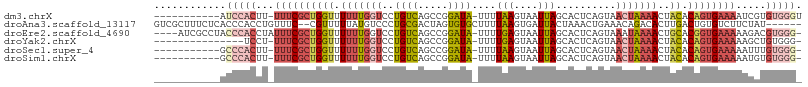
>dm3.chrX 5486837 95 - 22422827 -----------AUCCACUU-UUUCGCUGGUUUUUUGGUCCUGUCAGCCGGAUA-UUUUAAGUAAUUAGCACUCAGUAACUAAAACUACACAGUGAAAAUCGUGUGGGU -----------((((((.(-((((((((((.(((((((.(((...((..(.((-(.....))).)..))...)))..)))))))..)).)))))))))....)))))) ( -27.40, z-score = -3.14, R) >droAna3.scaffold_13117 4966616 100 + 5790199 GUCGCUUUCUCACCCACCUGUUUC--CGUUUUUAUGUCCCUGCGACUAGUGUGCUUUUAAGUGAUUACUAAACUGAAACAGACACUUGACUGUGUCUUCUAU------ ..................((((((--.((((....(((.((((.((....)))).....)).)))....)))).))))))(((((......)))))......------ ( -18.30, z-score = -1.33, R) >droEre2.scaffold_4690 2837978 102 - 18748788 ----AUCGCCUACCCACCUAUUUCGCUGGUUUUUUGGUCCUGUCAGCCGGAUA-UUUUGAGUAAUUAGCACUCAGUAAAUAAAACUGCACGGUGAAAAAGACGUGGG- ----........(((((((.((((((((((.(((((((((.(....).)))).-..((((((.......))))))....)))))..)).)))))))).))..)))))- ( -27.50, z-score = -1.50, R) >droYak2.chrX 18789007 90 + 21770863 ---------------UCCU-UUUCGCUGGUUUUUUGGUCCUGUCAGCCGGAUA-UUUUGAGUAAUUAGCACUCAGUAACUAAAACUACACAGUGAAAAAGCUGUGGG- ---------------...(-((((((((((.(((((((((.(....).)))..-..((((((.......))))))...))))))..)).))))))))).........- ( -21.10, z-score = -0.70, R) >droSec1.super_4 4908065 94 + 6179234 -----------GCCCACUU-UUUCGCUGGUUUUUUGGUCCUGUCAGCCGGAUA-UUUUAAGUAAUUAGCACUCAGUAACUAAAACUACACAGUGAAAAAUUUGUGGG- -----------.(((((((-((((((((((.(((((((.(((...((..(.((-(.....))).)..))...)))..)))))))..)).))))))))))...)))))- ( -28.30, z-score = -3.37, R) >droSim1.chrX 4249984 94 - 17042790 -----------GCCCACUU-UUUCGCUGGUUUUUUGGUCCUGUCAGCCGGAUA-UUUUAAGUAAUUAGCACUCAGUAACUAAAACUACACAGUGAAAAAUGUGUGGG- -----------.(((((((-((((((((((.(((((((.(((...((..(.((-(.....))).)..))...)))..)))))))..)).))))))))))...)))))- ( -28.30, z-score = -3.04, R) >consensus ___________ACCCACCU_UUUCGCUGGUUUUUUGGUCCUGUCAGCCGGAUA_UUUUAAGUAAUUAGCACUCAGUAACUAAAACUACACAGUGAAAAAUAUGUGGG_ ............(((((...((((((((....((((((.......))))))......(((....)))......................)))))))).....))))). (-10.85 = -13.02 + 2.17)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:18:04 2011