
| Sequence ID | dm3.chrX |
|---|---|
| Location | 5,408,968 – 5,409,068 |
| Length | 100 |
| Max. P | 0.614985 |

| Location | 5,408,968 – 5,409,068 |
|---|---|
| Length | 100 |
| Sequences | 7 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 74.17 |
| Shannon entropy | 0.44845 |
| G+C content | 0.49300 |
| Mean single sequence MFE | -31.23 |
| Consensus MFE | -12.59 |
| Energy contribution | -14.80 |
| Covariance contribution | 2.21 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.614985 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 5408968 100 - 22422827 GAAGUUCGAAAGAAAGC-----GGCAAGUUGGGCCGUAGCCAUUUCUCAACCUC----GACAAAGAAGUG--UGCCAGCUGGGUAUCUGUCAGAUACAGAUACUAAGUUGC ....(((....))).((-----(((.......))))).(((((((((.......----.....)))))))--.))(((((.(((((((((.....))))))))).))))). ( -34.10, z-score = -3.11, R) >droPer1.super_15 1200756 103 + 2181545 GGCGAUAGGAAGAGUACGAGAGAGUCAGUUUUGCCAGAGACGGGCAUCGAAGACAUAAAACAACAAAGCAACUGGCAGCUUGGUAUCUGUCAAAUACACGCAC-------- .(((...(........)((.((((((..(..((((.......))))..)..)))........((.((((........)))).)).))).)).......)))..-------- ( -20.70, z-score = -0.11, R) >dp4.chrXL_group1a 566985 101 + 9151740 GGCGAUAGGAAGAGUAC--GAGAGUCAGUUUUGCCAGAGACGGGCAUCGAAGACAUAAAACAACAAAGCAACUGGCAGCUUGGUAUCUGUCAAAUACACGCAC-------- .(((...((((((.(((--....)).).)))).))...(((((..((((....)...........((((........)))))))..))))).......)))..-------- ( -20.70, z-score = -0.18, R) >droSim1.chrX 4177092 100 - 17042790 GAAGUUCGGAAGAAAGC-----GGCAAGUUGGGCCGGAGCCAUUUCUCAACCUC----GACAAAGAAGUG--UGCCAGCUGGGUAUCUGUCAGAUACAGAUACUGAGUUGC ....(((....)))..(-----(((.......))))..(((((((((.......----.....)))))))--.))(((((.(((((((((.....))))))))).))))). ( -34.70, z-score = -2.29, R) >droSec1.super_4 4825674 100 + 6179234 GAAGUUCGGAAGAAAGC-----GGCAAGUUGGGCCGGAGCCAUUUCUCACCCUC----GACAAAGAAGUG--UGCCAGCUGGGUAUCUGUCAGAUACAGAUACUGAGUUGC ....(((....)))..(-----(((.......))))..(((((((((.......----.....)))))))--.))(((((.(((((((((.....))))))))).))))). ( -34.70, z-score = -2.03, R) >droYak2.chrX 18713070 100 + 21770863 GAAGUUCGAAAGAAAGC-----GGCAAGUUGGGCCGGAGCCAUUUCUCAACCUC----AACAUAGAAGUG--CGCCAGCUGGGUAUCUGUCAGAUACAGAUACUCAGUUGC ....(((....)))..(-----(((.......))))..(((((((((.......----.....)))))))--.))(((((((((((((((.....))))))))))))))). ( -37.70, z-score = -3.78, R) >droEre2.scaffold_4690 2762173 100 - 18748788 GAAGUUCGAAAGAAAGC-----GGCAAGUUGUGCCGGAGCCAUUUCUCAACCUC----GACAAAGAAGUG--UGCCAGCUGGGUAUCUGUCAGAUACAGAUACUGAGUUGC ....(((....)))..(-----((((.....)))))..(((((((((.......----.....)))))))--.))(((((.(((((((((.....))))))))).))))). ( -36.00, z-score = -3.31, R) >consensus GAAGUUCGGAAGAAAGC_____GGCAAGUUGGGCCGGAGCCAUUUCUCAACCUC____GACAAAGAAGUG__UGCCAGCUGGGUAUCUGUCAGAUACAGAUACUGAGUUGC (((.(((....)))........(((.......)))........))).............................(((((.(((((((((.....))))))))).))))). (-12.59 = -14.80 + 2.21)

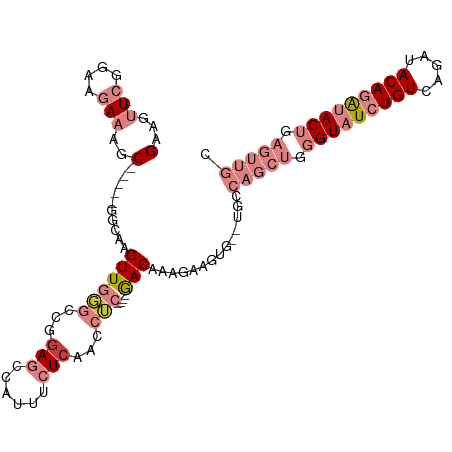

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:17:56 2011