
| Sequence ID | dm3.chrX |
|---|---|
| Location | 5,333,470 – 5,333,571 |
| Length | 101 |
| Max. P | 0.987240 |

| Location | 5,333,470 – 5,333,571 |
|---|---|
| Length | 101 |
| Sequences | 3 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 74.57 |
| Shannon entropy | 0.34531 |
| G+C content | 0.56389 |
| Mean single sequence MFE | -36.93 |
| Consensus MFE | -21.70 |
| Energy contribution | -23.57 |
| Covariance contribution | 1.87 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.91 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.27 |
| SVM RNA-class probability | 0.987240 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 5333470 101 + 22422827 GGGGGGGGCGGGGCG-GAGUGCUGUGAUUGAUGUUUAGCAUUUGACCCGGCGACUGUAGAUUGCAGUUUGCAGAUUGCCAUCCGCUCUCUCACUCACAAGAG .((((((((((((..-((((((((.(((....)))))))))))..)))(((((((((((((....)))))))).)))))...))))))))).(((....))) ( -45.60, z-score = -3.68, R) >droYak2.chrX 18635365 87 - 21770863 GGAGGGAGUAGGGGG-GAGUGCAGUGAUUGAUGUUUAGCAUUUGACCCAGGAGC----GACUGCAGAUUGCAGAUUGCCUGCCGCUCUC-CAC--------- ((((((....(((..-((((((...(((....)))..))))))..))).((.((----((((((.....)))).))))...)).)))))-)..--------- ( -35.10, z-score = -2.67, R) >droEre2.scaffold_4690 2687271 78 + 18748788 -------GCAGGGAGCGAGUGCUGUGAUUGAUGUUUAGCAUUUGACCCGG---C----GACUGCAGAUUGCAGAUUGCCUGCCGCUCUC-CAC--------- -------...((((((((((((((.(((....))))))))))......((---(----((((((.....)))).)))))...))))).)-)..--------- ( -30.10, z-score = -2.38, R) >consensus GG_GGG_GCAGGGAG_GAGUGCUGUGAUUGAUGUUUAGCAUUUGACCCGG___C____GACUGCAGAUUGCAGAUUGCCUGCCGCUCUC_CAC_________ .((((((((.(((...((((((((.(((....)))))))))))..)))............((((.....))))..........))))))))........... (-21.70 = -23.57 + 1.87)

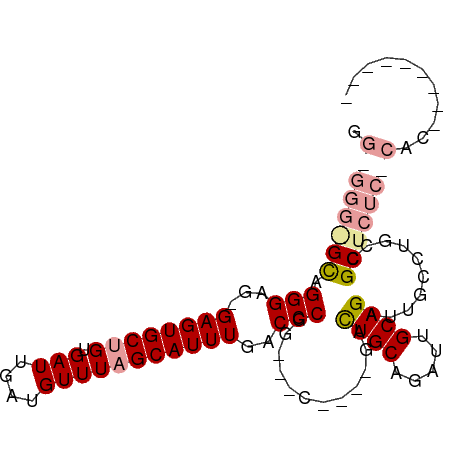
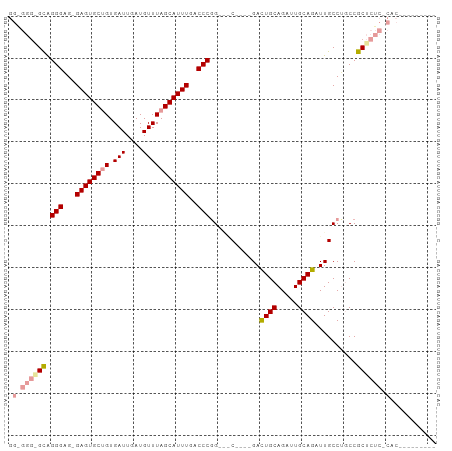
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:17:45 2011