
| Sequence ID | dm3.chrX |
|---|---|
| Location | 5,280,522 – 5,280,625 |
| Length | 103 |
| Max. P | 0.935424 |

| Location | 5,280,522 – 5,280,623 |
|---|---|
| Length | 101 |
| Sequences | 3 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 89.00 |
| Shannon entropy | 0.15207 |
| G+C content | 0.37491 |
| Mean single sequence MFE | -25.33 |
| Consensus MFE | -22.17 |
| Energy contribution | -21.07 |
| Covariance contribution | -1.10 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.87 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.40 |
| SVM RNA-class probability | 0.935424 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
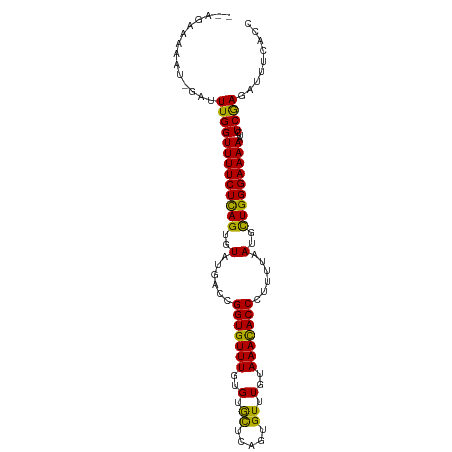
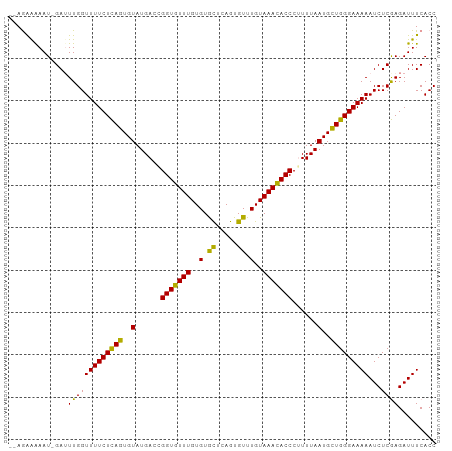
>dm3.chrX 5280522 101 - 22422827 AAAAAAAUGGAGUUGGUUUUCUCAGUGUAUGACCGGUGUUUGUGUGCUCAGUGUUUGUAAAUACCCUUUUAAUGCUGGGAAAAAUCUCGAGAUUUCACCUG .......(((((((..((((((((((((......(((((((..(.((.....)).)..)))))))......)))))))))))).......))))))).... ( -24.70, z-score = -1.97, R) >droSec1.super_4 4708033 97 + 6179234 --AAAAAU-GAUUUGGUUUUCUCAGUGUAUGA-GGGUGUUUGUGUGCUCAGUGUUUGUAAACACCCUUUUAAUGCUGGGAAAAAUCUCGAGAUUUCACCUG --......-.....(((.((((((((((..((-((((((((..(.((.....)).)..))))))))))...))))))))))(((((....))))).))).. ( -31.30, z-score = -4.25, R) >droEre2.scaffold_4690 2637867 98 - 18748788 --GGGAAU-GAUUUGGUUUUCUUAGUGUAUGACUGGUGUUUGUGUAUUCAGUGUUUGUAAACACCCAUUUAAUGUUGGGAAAAAUCUCAAGAUUUCACCUG --(((((.-..((((((((((((((..(..((..(((((((..(.((.....)).)..)))))))..))..)..)))))))))...)))))..))).)).. ( -20.00, z-score = -0.17, R) >consensus __AAAAAU_GAUUUGGUUUUCUCAGUGUAUGACCGGUGUUUGUGUGCUCAGUGUUUGUAAACACCCUUUUAAUGCUGGGAAAAAUCUCGAGAUUUCACCUG ..............(((.(((((((..(.(((..(((((((..(.((.....)).)..)))))))...))))..)))))))(((((....))))).))).. (-22.17 = -21.07 + -1.10)



| Location | 5,280,524 – 5,280,625 |
|---|---|
| Length | 101 |
| Sequences | 3 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 88.33 |
| Shannon entropy | 0.16117 |
| G+C content | 0.37491 |
| Mean single sequence MFE | -26.10 |
| Consensus MFE | -21.33 |
| Energy contribution | -19.90 |
| Covariance contribution | -1.43 |
| Combinations/Pair | 1.27 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.13 |
| SVM RNA-class probability | 0.896459 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 5280524 101 - 22422827 AGAAAAAAAUGGAGUUGGUUUUCUCAGUGUAUGACCGGUGUUUGUGUGCUCAGUGUUUGUAAAUACCCUUUUAAUGCUGGGAAAAAUCUCGAGAUUUCACC .((((....(.(((....((((((((((((......(((((((..(.((.....)).)..)))))))......))))))))))))..))).)..))))... ( -26.70, z-score = -2.43, R) >droSec1.super_4 4708035 97 + 6179234 --AGAAAAAU-GAUUUGGUUUUCUCAGUGUAUGA-GGGUGUUUGUGUGCUCAGUGUUUGUAAACACCCUUUUAAUGCUGGGAAAAAUCUCGAGAUUUCACC --.((((..(-((.....((((((((((((..((-((((((((..(.((.....)).)..))))))))))...))))))))))))...)))...))))... ( -31.90, z-score = -4.21, R) >droEre2.scaffold_4690 2637869 98 - 18748788 --AGGGGAAU-GAUUUGGUUUUCUUAGUGUAUGACUGGUGUUUGUGUAUUCAGUGUUUGUAAACACCCAUUUAAUGUUGGGAAAAAUCUCAAGAUUUCACC --.(((((((-..((((((((((((((..(..((..(((((((..(.((.....)).)..)))))))..))..)..)))))))))...)))))))))).)) ( -19.70, z-score = 0.11, R) >consensus __AGAAAAAU_GAUUUGGUUUUCUCAGUGUAUGACCGGUGUUUGUGUGCUCAGUGUUUGUAAACACCCUUUUAAUGCUGGGAAAAAUCUCGAGAUUUCACC ...((((..(.((.....(((((((((..(.(((..(((((((..(.((.....)).)..)))))))...))))..)))))))))...)).)..))))... (-21.33 = -19.90 + -1.43)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:17:33 2011