
| Sequence ID | dm3.chrX |
|---|---|
| Location | 5,222,955 – 5,223,046 |
| Length | 91 |
| Max. P | 0.734126 |

| Location | 5,222,955 – 5,223,046 |
|---|---|
| Length | 91 |
| Sequences | 15 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 60.29 |
| Shannon entropy | 0.90084 |
| G+C content | 0.55202 |
| Mean single sequence MFE | -27.51 |
| Consensus MFE | -11.32 |
| Energy contribution | -11.38 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.50 |
| Mean z-score | -0.53 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.734126 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 5222955 91 - 22422827 UCCCGGACUCAAAUUCCGGCUGCAUGGGCGACACCAAGUUCCACAUGCGAUUGCGC----UGCACAACGCGUCG---GGAUUCCUUGCCGGGCUACAA------------- .(((((.....(((((((((((((((((.(((.....))))).))))))...(((.----.......)))))))---))))).....)))))......------------- ( -35.50, z-score = -2.34, R) >droSim1.chrX 4022682 91 - 17042790 UCCCGGACUCAAAUUCCGGCUGCAUGGGCGACACCAAGUUCCAUAUGCGAUUGCGC----UGCACAACGCGUCG---GGACUCGUUGCCGGGCUACAA------------- .(((((.......(((((((((((((((.(((.....))))).))))))...(((.----.......)))))))---))).......)))))......------------- ( -30.84, z-score = -0.58, R) >droSec1.super_4 4652107 91 + 6179234 UCCCGGACUCAAAUUCCGGCUGCAUGGGCGACACCAAGUUCCAUAUGCGAUUGCGC----UGCACAACGCGUCG---GGACUCGUUGCCGGGCUACAA------------- .(((((.......(((((((((((((((.(((.....))))).))))))...(((.----.......)))))))---))).......)))))......------------- ( -30.84, z-score = -0.58, R) >droYak2.chrX 18527792 91 + 21770863 UUCCGGACUCCAAUUCCGGCUGCAUGGGCGACACCAAGUUCCACAUGCGAUUGCGC----UGCACAACGCGUCG---UGAUUCCCUGCCAGGCUACAA------------- ..(((((.......)))(((.(((((((.(((.....))))).)))))(((..((.----(((.....))).))---..)))....))).))......------------- ( -26.90, z-score = -0.26, R) >droEre2.scaffold_4690 2578340 91 - 18748788 UUCCGGACUCCAAUUCCGGCUGCAUGGGCGACACCAAGUUCCACAUGCGACUGCGC----UGCACAACGCGACG---GGAUUCCUUGCCAGGCUACAA------------- ..(((((.......)))(((((((((((.(((.....))))).)))))).((((((----........))).))---)........))).))......------------- ( -25.30, z-score = 0.22, R) >droAna3.scaffold_13117 2668837 91 - 5790199 UCCCGGACUCCAACUCCGGCUGCAUGGGCGACACCAAGUUCCACAUGCGCCUGCGA----UGCACUCGCAGACA---GGAAUCCUUGCCAGGAUACAA------------- (((((((.......))))((.(((((((.(((.....))))).)))))))((((((----.....))))))...---)))(((((....)))))....------------- ( -29.80, z-score = -1.49, R) >dp4.chrXL_group1e 10809096 109 - 12523060 UCCCCGACUCCAACUCGGGCUGCAUGGGCGACACCAAGUUCCACAUGCGUCUGCGUGCCAUCUCCA-AGCAGCAGCAACAGCACCAGCAGAAGCCCCUGC-CCACCUACAA ...((((.......))))((((((((((.(((.....))))).)))))(.((((.(((........-.))))))))...)))....((((......))))-.......... ( -29.70, z-score = -0.37, R) >droPer1.super_14 253868 109 - 2168203 UCCCCGACUCCAACUCGGGCUGCAUGGGCGACACCAAGUUCCACAUGCGUCUGCGUGCCAUCUCCA-AGCAGCAGCAACAGCACCAACAGAAGCCCCUGC-CCACCUACAA ..(((((.......)))))..(((.(((((((.....))).....(((((((((.(((........-.)))))))..)).))).........)))).)))-.......... ( -27.50, z-score = -0.21, R) >droWil1.scaffold_181150 4359746 100 - 4952429 UCCCCGACUCGAAUUCGGGCUGUAUGGGCGACACCAAAUUCCAUAUGCGUUUGCGUGCCCUGUCCA-AAC-------CCAGAGCCUAU-GCGCCCAUUAC-CCA-AUACAA ................((((.((((((((..((((((((.(.....).))))).)))..(((....-...-------.))).))))))-)))))).....-...-...... ( -27.40, z-score = -1.98, R) >droVir3.scaffold_12932 1726285 106 + 2102469 UCCCGGACUCCAAUUCGGGCUGCAUGGGCGACACCAAGUUCCAUAUGCGUCUGCGGGCCAUCCAAAAUGCCAAACAUAAAAUGCCCGCCACCAUGAAC-----CAAUACAA .((((((......))))))..(((((((.(((.....))))).)))))....((((((........(((.....))).....))))))..........-----........ ( -25.32, z-score = 0.33, R) >droMoj3.scaffold_6308 182166 106 - 3356042 UCCCAGACUCCAAUUCUGGCUGUAUGGGCGACACCAAGUUCCAUAUGCGCCUGCGAGCGUUUCAACAUACAAAACAUGAAAUGCCCGCCAAUUUGCA-----AAAUUACAA ..(((((.......))))).((((.(((((.((............)))))))(((.((((((((............)))))))).))).....))))-----......... ( -25.70, z-score = -1.54, R) >droGri2.scaffold_14853 2423422 106 - 10151454 UUCCCGACUCCAAUUCCGGUUGCAUGGGCGACACCAAGUUCCAUAUGCGUCUCCGUGCGCUAAACACCGCAAAACACAAAACACCGACAACGAUGCU-----ACACUACAA ................((((.(((((((.(((.(............).))))))))))((........))............))))...........-----......... ( -18.00, z-score = 0.45, R) >anoGam1.chr2L 48425534 109 - 48795086 UCCCGGAUUCGAAUUCCGGGUGCAUGGGAGACUCGCAGUUCCACAUCCGGUUGCGUGUGUCGUCCGGCUCGGAAAACUCCAUACUCCCCAAGGGGCUGC--AGGAGUUCAA .((((((.......)))))).((.(((..(((.(((((..((......)))))))...)))..)))))......((((((.....(((....)))....--.))))))... ( -39.60, z-score = -0.51, R) >apiMel3.Group2 8536207 103 + 14465785 UUCCUGACUCUAACUCUGGUUGCAUGGGAGAUACACAAUAUCAUGUCAGAAUAAGA---CAAAAUGGAGCUAUACUUCAAGAGACUAAAGCCUUAAA-----AGAAUAUGA .......((((...((((...((((((.............))))))))))......---.....(((((.....)))))))))..............-----......... ( -13.92, z-score = 1.30, R) >triCas2.chrUn_132 216126 103 + 222081 UCCCCGACUCCAAUUCGGGAUGUAUGGGGGACACCACCUU-------CAUCAUCAGGCU-UCCAAACAGCCCAGGAAAAAACACGCGCCGGGAUGAACACUUGCAAUACAA ..(((((.......))))).((((((((((......))))-------).(((((.((((-(((..........)))).........)))..))))).........))))). ( -26.30, z-score = -0.42, R) >consensus UCCCGGACUCCAAUUCCGGCUGCAUGGGCGACACCAAGUUCCACAUGCGACUGCGCG___UGCACAACGCGACA___GGAAUCCCUGCCGGGCUACAA________UACAA ..(((((.......))))).((((((((.(((.....))))).)))))).............................................................. (-11.32 = -11.38 + 0.06)


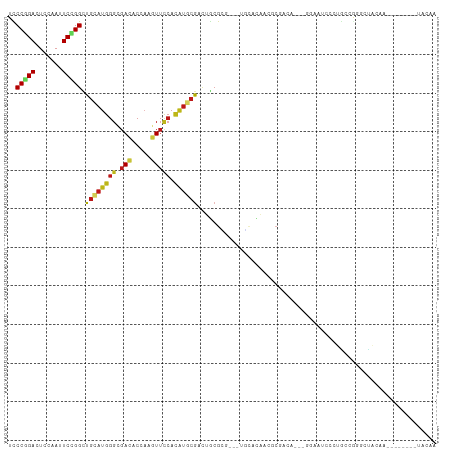
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:17:16 2011