
| Sequence ID | dm3.chrX |
|---|---|
| Location | 4,921,298 – 4,921,351 |
| Length | 53 |
| Max. P | 0.957292 |

| Location | 4,921,298 – 4,921,351 |
|---|---|
| Length | 53 |
| Sequences | 3 |
| Columns | 68 |
| Reading direction | reverse |
| Mean pairwise identity | 61.38 |
| Shannon entropy | 0.50206 |
| G+C content | 0.60137 |
| Mean single sequence MFE | -8.93 |
| Consensus MFE | -6.43 |
| Energy contribution | -6.77 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.72 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.64 |
| SVM RNA-class probability | 0.957292 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
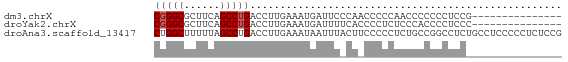
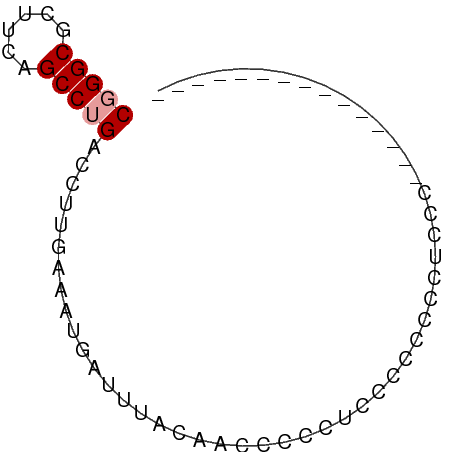
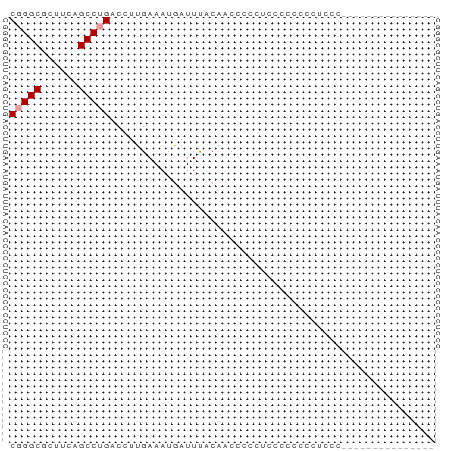
>dm3.chrX 4921298 53 - 22422827 CGGGCGCUUCAGCCUGACCUUGAAAUGAUUCCCAACCCCCAACCCCCCCUCCG--------------- (((((......))))).....................................--------------- ( -8.10, z-score = -1.11, R) >droYak2.chrX 18229837 53 + 21770863 CGGGCGCUUCAGCCUGACCUUGAAAUGAUUUUCACCCCUCUCCCACCCCUCCC--------------- (((((......)))))....(((((....)))))...................--------------- ( -10.10, z-score = -1.85, R) >droAna3.scaffold_13417 276836 68 - 6960332 CUGGCUUUUUAGCCUGACCUUGAAAUAAUUUACUUCCCCCUCUGCCGGCCUCUGCCUCCCCCUCUCCG ..((((....))))................................(((....)))............ ( -8.60, z-score = -1.12, R) >consensus CGGGCGCUUCAGCCUGACCUUGAAAUGAUUUACAACCCCCUCCCCCCCCUCCC_______________ (((((......))))).................................................... ( -6.43 = -6.77 + 0.33)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:16:44 2011