
| Sequence ID | dm3.chrX |
|---|---|
| Location | 4,889,572 – 4,889,651 |
| Length | 79 |
| Max. P | 0.831622 |

| Location | 4,889,572 – 4,889,651 |
|---|---|
| Length | 79 |
| Sequences | 4 |
| Columns | 90 |
| Reading direction | reverse |
| Mean pairwise identity | 56.61 |
| Shannon entropy | 0.67661 |
| G+C content | 0.47567 |
| Mean single sequence MFE | -20.95 |
| Consensus MFE | -10.88 |
| Energy contribution | -9.82 |
| Covariance contribution | -1.06 |
| Combinations/Pair | 1.44 |
| Mean z-score | -0.76 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.84 |
| SVM RNA-class probability | 0.831622 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
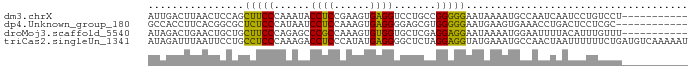
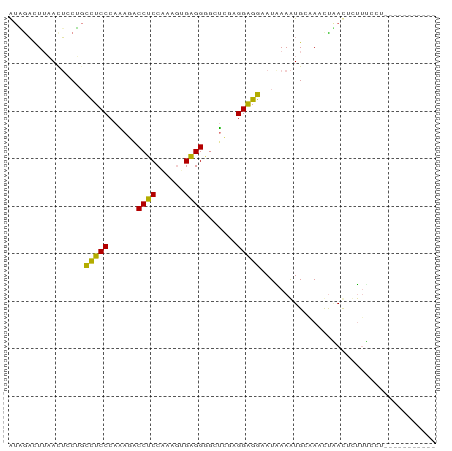
>dm3.chrX 4889572 79 - 22422827 AUUGACUUAACUCCAGCUUCCCAAAUACCUCCGAAGUGAGGUCCUGCCGGGGGAAUAAAAUGCCAAUCAAUCCUGUCCU----------- (((((............(((((....(((((......)))))((....)))))))...........)))))........----------- ( -13.50, z-score = 0.91, R) >dp4.Unknown_group_180 11568 78 - 35195 GCCACCUUCACGGCGCUCUCCCAUAAUCCUCCAAAGUGAGGGGAGCGUGGGGGAAUGAAGUGAAACCUGACUCCUCGC------------ ......(((((..((.((((((((..((((((.....).)))))..)))))))).))..)))))..............------------ ( -27.80, z-score = -1.27, R) >droMoj3.scaffold_5540 3978 79 + 4752 AUAGACUGAACUGCUGCUUCCCAGAGCCCGCCAAAGUGUGGUGCUCGAGGAGGAAUAAAAUGGAAUUUUACAUUUGUUU----------- .......((((.....(((((..((((((((......)))).))))..)))))....(((((........)))))))))----------- ( -22.50, z-score = -1.97, R) >triCas2.singleUn_1341 16365 90 + 21755 AUAGAUUUAAUUCCUGCCUCCCAAAGACCUCCCAUAUGAGGGGCUCUAGGAGGUAUGAAAUGCCAACUAAUUUUUUCUGAUGUCAAAAAU ..........(((.(((((((...(((.(((((....).)))).))).))))))).)))............................... ( -20.00, z-score = -0.69, R) >consensus AUAGACUUAACUCCUGCCUCCCAAAGACCUCCAAAGUGAGGGGCUCGAGGAGGAAUAAAAUGCAAACUAACUCUUUCCU___________ ................(((((......((((......)))).......)))))..................................... (-10.88 = -9.82 + -1.06)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:16:42 2011