
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 10,138,377 – 10,138,482 |
| Length | 105 |
| Max. P | 0.846329 |

| Location | 10,138,377 – 10,138,482 |
|---|---|
| Length | 105 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 51.51 |
| Shannon entropy | 0.76357 |
| G+C content | 0.56588 |
| Mean single sequence MFE | -35.30 |
| Consensus MFE | -11.98 |
| Energy contribution | -11.48 |
| Covariance contribution | -0.50 |
| Combinations/Pair | 1.47 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.34 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.89 |
| SVM RNA-class probability | 0.846329 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
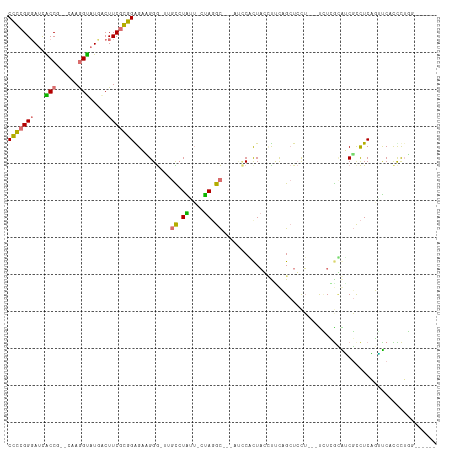
>dm3.chr2L 10138377 105 + 23011544 CACCGGGACUGCCAGUCAAGGUAUGACUUCGGUGAGCAGGA----UGUGAGGCCAGGUCGCACUCCCUUCCUUCAG-UGUAGUCCAUGGGGGCUCCUCCUUCCACCCUUU------ .(((..(((.....)))..)))........((((.(.((((----.(.(((.((.((.((((((..........))-))).).))....)).)))))))).)))))....------ ( -31.90, z-score = 1.49, R) >dp4.Unknown_group_238 12184 97 - 16630 CCGCGGGAUCACCG--UUAGGUAUUGCUUCGCGGGGAAGGG-GGGCCUAUU-CUAGGC------CGCUACAUCCCGCUCUU---UCUCGCAUCGUCUCAGGUCAUCCUGG------ ..((((((..(((.--...)))........((((((...((-.((((((..-.)))))------).))...))))))....---)))))).......((((....)))).------ ( -41.00, z-score = -2.59, R) >droVir3.scaffold_13324 1668592 103 - 2960039 CCGCGGGAUCACCG--CAAAGUAUUACUUCGCGGGGAUGGG-UUGCCUAUU-UUAGGCAGUAUCCAUUAGUCGUCGCUCCU---UUUCGCAUCGUCUCAGAUCCCUCUAG------ ....((((((.(((--(.(((.....))).)))).((((((-(((((((..-.))))))..)))))))((.((..((....---....))..)).))..)))))).....------ ( -37.70, z-score = -4.00, R) >triCas2.chrUn_190 47221 111 - 65089 CCCCGGUAUCGCCG--UUAGGCAUGUCUUCAGGGAGCCGGGCUUGUCUGUUUCUACGC---CUCCACUAGCUUCAGCUCCUGCGUCUAAUGACGCUUCUUUUCAUUUAGCCAUUGC ...(((.....)))--...(((.........((((((.(((((.((..((......))---....)).)))))..))))))(((((....))))).............)))..... ( -30.60, z-score = -0.63, R) >consensus CCCCGGGAUCACCG__CAAGGUAUGACUUCGCGGAGAAGGG_UUGCCUAUU_CUAGGC___AUCCACUACCUUCAGCUCCU___UCUCGCAUCGCCUCAGUUCACCCUGG______ (((((((...(((......))).....)))))))..........(((((....))))).......................................................... (-11.98 = -11.48 + -0.50)


| Location | 10,138,377 – 10,138,482 |
|---|---|
| Length | 105 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 51.51 |
| Shannon entropy | 0.76357 |
| G+C content | 0.56588 |
| Mean single sequence MFE | -33.67 |
| Consensus MFE | -10.41 |
| Energy contribution | -10.23 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.41 |
| Structure conservation index | 0.31 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.705414 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 10138377 105 - 23011544 ------AAAGGGUGGAAGGAGGAGCCCCCAUGGACUACA-CUGAAGGAAGGGAGUGCGACCUGGCCUCACA----UCCUGCUCACCGAAGUCAUACCUUGACUGGCAGUCCCGGUG ------..((((((..((((((.((((((.....((...-....))...))).).))..)))..)))..))----))))...(((((.(((((.....)))))((....))))))) ( -31.70, z-score = 1.36, R) >dp4.Unknown_group_238 12184 97 + 16630 ------CCAGGAUGACCUGAGACGAUGCGAGA---AAGAGCGGGAUGUAGCG------GCCUAG-AAUAGGCCC-CCCUUCCCCGCGAAGCAAUACCUAA--CGGUGAUCCCGCGG ------.((((....))))......((((.((---....(((((..(..(.(------(((((.-..)))))))-..)...))))).......((((...--.)))).)).)))). ( -34.60, z-score = -2.58, R) >droVir3.scaffold_13324 1668592 103 + 2960039 ------CUAGAGGGAUCUGAGACGAUGCGAAA---AGGAGCGACGACUAAUGGAUACUGCCUAA-AAUAGGCAA-CCCAUCCCCGCGAAGUAAUACUUUG--CGGUGAUCCCGCGG ------...(.((((((..((.((.(((....---....))).)).))...((((..((((((.-..)))))).-...))))((((((((.....)))))--))).)))))).).. ( -36.50, z-score = -3.81, R) >triCas2.chrUn_190 47221 111 + 65089 GCAAUGGCUAAAUGAAAAGAAGCGUCAUUAGACGCAGGAGCUGAAGCUAGUGGAG---GCGUAGAAACAGACAAGCCCGGCUCCCUGAAGACAUGCCUAA--CGGCGAUACCGGGG ((...((((............(((((....)))))....((((...(((((....---)).)))...))).).))))..)).(((((......((((...--.))))....))))) ( -31.90, z-score = -0.60, R) >consensus ______CCAGAAUGAAAUGAGACGACGCGAGA___AAGAGCGGAAGCUAGUGGAU___GCCUAG_AACAGACAA_CCCUGCCCCCCGAAGUAAUACCUAA__CGGCGAUCCCGCGG ..........................................................(((((....)))))..........(((((......((((......))))....))))) (-10.41 = -10.23 + -0.19)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:29:44 2011