
| Sequence ID | dm3.chrX |
|---|---|
| Location | 4,883,581 – 4,883,675 |
| Length | 94 |
| Max. P | 0.688341 |

| Location | 4,883,581 – 4,883,675 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 85.38 |
| Shannon entropy | 0.25581 |
| G+C content | 0.48566 |
| Mean single sequence MFE | -29.25 |
| Consensus MFE | -18.19 |
| Energy contribution | -19.72 |
| Covariance contribution | 1.53 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.62 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.623475 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 4883581 94 + 22422827 AUCUGCCAAGCGU-UUUCUUCCGGAAAAGUCAAUCAGUCCAUCACUCAAUCGACGAUUGA-------AGGAAGCCAAACAGUUGGCAG--UCUGCAGACUGCCG------ ........(((((-((.(((((.......((((((.(((.((......)).)))))))))-------.)))))..)))).)))(((((--((....))))))).------ ( -28.30, z-score = -2.37, R) >droSim1.chrX 3784600 96 + 17042790 AUCUGCCGAGCGU-UUUCUUCCGGAAAAGUCAAUCAGUCCAUCACUCAAUCGACGAUUGA-------AGGAAGCCAAACAGUUGGCAGAGUCUGCAGACUGCCG------ .((((((((..((-((.(((((.......((((((.(((.((......)).)))))))))-------.)))))..))))..))))))))(((....))).....------ ( -29.80, z-score = -2.00, R) >droSec1.super_4 4323381 96 - 6179234 AUCUGCCGAGCGU-UUUCUUCCGGAAAAGUCAAUCAGUCCAUCACUCAAUCGACGAUUGA-------AGGAAGCCAAACAGUUGGCAGAGUCUGCAGACUGCCG------ .((((((((..((-((.(((((.......((((((.(((.((......)).)))))))))-------.)))))..))))..))))))))(((....))).....------ ( -29.80, z-score = -2.00, R) >droYak2.chrX 18199477 101 - 21770863 AUCUGCCGAGUGU-UUUCUUCCGGAAAAGUCAAUCAGUCCAUCACUCAAUCGACGAUUGAGGAUUGAAGGAAGCCAAACAGUUGGCAG--UCUGCAGACUGCCG------ .......((.(((-((.(((((.((....))..(((((((.(((.((.......)).)))))))))).)))))..))))).))(((((--((....))))))).------ ( -35.30, z-score = -2.99, R) >droEre2.scaffold_4690 2249010 100 + 18748788 AUCUGCCGCGUGA-UUUCUUCCGGAAAAGUCAAUCAGUCCAUCACUCAAUCGACGAUUGA-------AGGAAGCCAAACAGUUGGCAG--UCUACAGACUGCAACUGCCG .......((((..-...(((((.......((((((.(((.((......)).)))))))))-------.)))))....))(((((.(((--((....)))))))))))).. ( -27.40, z-score = -1.52, R) >droAna3.scaffold_13417 240934 89 + 6960332 AUCGGACGAGUGGCUUUCUUCCGGAAAAGUCAAUCACUCAAUCAUUCAAUCAUUGACUGA-------AGGAAGCCAAA--ACUG------UUUGCAGACUGCCG------ ..(((.(.(((((((((((((......(((((((.................)))))))))-------))))))))...--.(((------....))))))))))------ ( -24.93, z-score = -1.96, R) >consensus AUCUGCCGAGCGU_UUUCUUCCGGAAAAGUCAAUCAGUCCAUCACUCAAUCGACGAUUGA_______AGGAAGCCAAACAGUUGGCAG__UCUGCAGACUGCCG______ ..(((((((........(((((.......((((((.(((.((......)).)))))))))........)))))........)))))))...................... (-18.19 = -19.72 + 1.53)

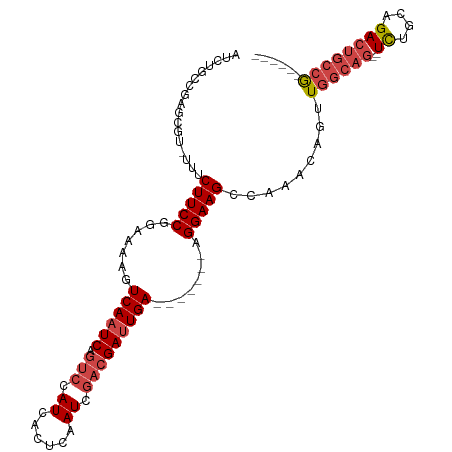
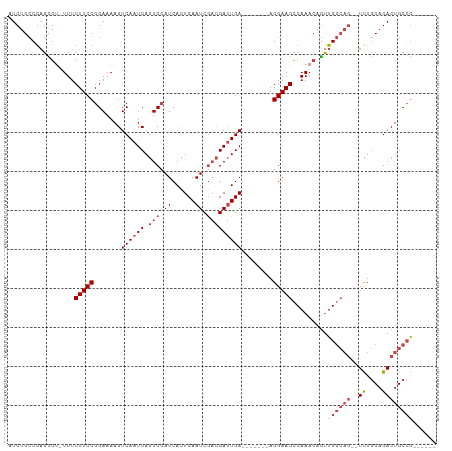
| Location | 4,883,581 – 4,883,675 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 85.38 |
| Shannon entropy | 0.25581 |
| G+C content | 0.48566 |
| Mean single sequence MFE | -31.00 |
| Consensus MFE | -22.66 |
| Energy contribution | -23.74 |
| Covariance contribution | 1.09 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.73 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.688341 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 4883581 94 - 22422827 ------CGGCAGUCUGCAGA--CUGCCAACUGUUUGGCUUCCU-------UCAAUCGUCGAUUGAGUGAUGGACUGAUUGACUUUUCCGGAAGAAA-ACGCUUGGCAGAU ------.(((((((....))--)))))..((((..(((.....-------(((((((((.(((....))).))).))))))......(....)...-..)))..)))).. ( -35.20, z-score = -3.30, R) >droSim1.chrX 3784600 96 - 17042790 ------CGGCAGUCUGCAGACUCUGCCAACUGUUUGGCUUCCU-------UCAAUCGUCGAUUGAGUGAUGGACUGAUUGACUUUUCCGGAAGAAA-ACGCUCGGCAGAU ------.....(((....)))((((((...(((((..(((((.-------(((((((((.(((....))).))).)))))).......))))).))-)))...)))))). ( -30.40, z-score = -1.33, R) >droSec1.super_4 4323381 96 + 6179234 ------CGGCAGUCUGCAGACUCUGCCAACUGUUUGGCUUCCU-------UCAAUCGUCGAUUGAGUGAUGGACUGAUUGACUUUUCCGGAAGAAA-ACGCUCGGCAGAU ------.....(((....)))((((((...(((((..(((((.-------(((((((((.(((....))).))).)))))).......))))).))-)))...)))))). ( -30.40, z-score = -1.33, R) >droYak2.chrX 18199477 101 + 21770863 ------CGGCAGUCUGCAGA--CUGCCAACUGUUUGGCUUCCUUCAAUCCUCAAUCGUCGAUUGAGUGAUGGACUGAUUGACUUUUCCGGAAGAAA-ACACUCGGCAGAU ------.(((((((....))--)))))..(((((((.(((((....((((((((((...))))))).)))(((............))))))))...-.))...))))).. ( -34.50, z-score = -2.63, R) >droEre2.scaffold_4690 2249010 100 - 18748788 CGGCAGUUGCAGUCUGUAGA--CUGCCAACUGUUUGGCUUCCU-------UCAAUCGUCGAUUGAGUGAUGGACUGAUUGACUUUUCCGGAAGAAA-UCACGCGGCAGAU ((((((((((((((....))--))).)))))))....(((((.-------(((((((((.(((....))).))).)))))).......)))))...-.....))...... ( -31.60, z-score = -1.26, R) >droAna3.scaffold_13417 240934 89 - 6960332 ------CGGCAGUCUGCAAA------CAGU--UUUGGCUUCCU-------UCAGUCAAUGAUUGAAUGAUUGAGUGAUUGACUUUUCCGGAAGAAAGCCACUCGUCCGAU ------((((...(((....------))).--..((((((.((-------(((((((((.(((.(.....).))).)))))))......)))).))))))...).))).. ( -23.90, z-score = -0.89, R) >consensus ______CGGCAGUCUGCAGA__CUGCCAACUGUUUGGCUUCCU_______UCAAUCGUCGAUUGAGUGAUGGACUGAUUGACUUUUCCGGAAGAAA_ACACUCGGCAGAU ........((.....)).....(((((...(((.(..(((((........(((((((((.(((....))).))).)))))).......)))))..).)))...))))).. (-22.66 = -23.74 + 1.09)


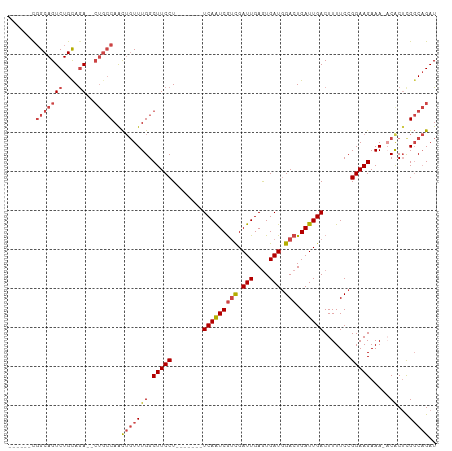
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:16:41 2011