
| Sequence ID | dm3.chrX |
|---|---|
| Location | 4,868,002 – 4,868,093 |
| Length | 91 |
| Max. P | 0.682767 |

| Location | 4,868,002 – 4,868,093 |
|---|---|
| Length | 91 |
| Sequences | 7 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 64.30 |
| Shannon entropy | 0.64888 |
| G+C content | 0.37523 |
| Mean single sequence MFE | -21.51 |
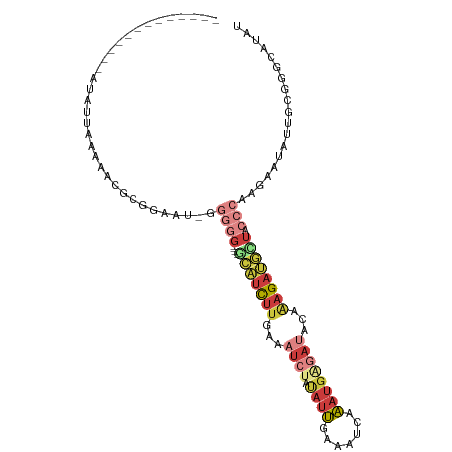
| Consensus MFE | -5.57 |
| Energy contribution | -7.37 |
| Covariance contribution | 1.80 |
| Combinations/Pair | 1.50 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.26 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.682767 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
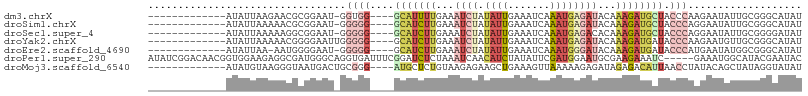
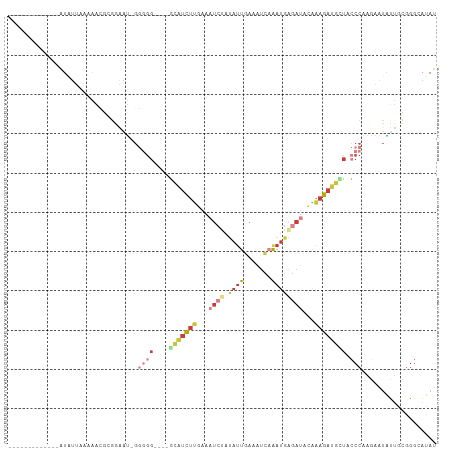
>dm3.chrX 4868002 91 + 22422827 -------------AUAUUAAGAACGCGGAAU-GGUGG----GCAUUUUGAAAUCUAUAUUGAAAUCAAAUGAGAUACAAAGAUGCUACCCAAGAAUAUUGCGGGCAUAU -------------.......(..(((((..(-((..(----(((((((...((((...(((....)))...))))...))))))))..)))......)))))..).... ( -22.30, z-score = -2.78, R) >droSim1.chrX 3768417 91 + 17042790 -------------AUAUUAAAAACGCGGAAU-GGGGG----GCAUCUUGAAAUCUAUAUUGAAAUCAAAUGAGAUACAAAGAUGCUACCCAGGAAUAUUGCGGGCAUAU -------------..........(((((..(-(((..----(((((((...((((...(((....)))...))))...)))))))..))))......)))))....... ( -27.70, z-score = -4.51, R) >droSec1.super_4 4308484 91 - 6179234 -------------AUAUUAAAAAGGCGGAAU-GGGGG----GCAUCUUGAAAUCUAUAUUGAAAUCAAAUGAGACACAAAGAUGCUACCCAGGAAUAUUGCGGGGAUAU -------------...........((((..(-(((..----(((((((....(((...(((....)))...)))....)))))))..))))......))))........ ( -23.80, z-score = -3.29, R) >droYak2.chrX 18183542 92 - 21770863 -------------AUAUUAAAAACGGGGAAUUGGGGG----GCAUCUUGAAAUCUAUAUUGAAAUCAAAUGAGAUACAAAGAUGAUACCCAAGAAUGUUGCGGGCAUAU -------------..........((.((..(((((..----.((((((...((((...(((....)))...))))...))))))...))))).....)).))....... ( -21.70, z-score = -2.62, R) >droEre2.scaffold_4690 2233340 90 + 18748788 -------------AUAUUAA-AAUGGGGAAU-GGGGG----GCAUCUUGAAAUCUAUAUUGAAAUCAAAUGGGAUACAAAGAUGAUACCCAUGAAUAUGGCGGGCAUAU -------------.......-........((-(((..----.((((((...((((...(((....)))...))))...))))))...)))))..(((((.....))))) ( -20.80, z-score = -2.04, R) >droPer1.super_290 1179 104 + 47041 AUAUCGGACAACGGUGGAAGAGGCGAUGGGCAGGUGAUUUCGGAUCUCUAAAUCAACAUCUAUAUUCGAUGGAAUGCGAAGAAAUC-----GAAAUGGCAUACGAAUAC .(((((.....))))).......((...(.((..(((((((.(((......)))..((((.......)))).........))))))-----)...)).)...))..... ( -18.50, z-score = 0.71, R) >droMoj3.scaffold_6540 32149132 92 + 34148556 -------------AUAUGUAAGGGUAAUGACUGCGGG----AUGCUCUGUAAGAGAAGCUGAAAGUUAAAAAGAGAUAGAGACAUUAACCUAUACAGCUAUAGGUAUAU -------------...((((.(........))))).(----(((((((((......(((.....)))........)))))).)))).(((((((....))))))).... ( -15.74, z-score = -0.28, R) >consensus _____________AUAUUAAAAACGCGGAAU_GGGGG____GCAUCUUGAAAUCUAUAUUGAAAUCAAAUGAGAUACAAAGAUGCUACCCAAGAAUAUUGCGGGCAUAU ..................................(((....(((((((...((((.((((.......))))))))...)))))))..)))................... ( -5.57 = -7.37 + 1.80)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:16:36 2011