
| Sequence ID | dm3.chrX |
|---|---|
| Location | 4,781,163 – 4,781,257 |
| Length | 94 |
| Max. P | 0.895959 |

| Location | 4,781,163 – 4,781,257 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 87.45 |
| Shannon entropy | 0.20938 |
| G+C content | 0.45058 |
| Mean single sequence MFE | -27.48 |
| Consensus MFE | -15.43 |
| Energy contribution | -15.27 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.81 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.796264 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
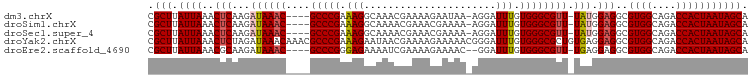
>dm3.chrX 4781163 94 + 22422827 CGCUUAUUAAACUCAAGAUAAAC----GCCCGAAAGGCAAACGAAAAGAAUAA-AGGAUUUGUGGGCGUU-UAUGGAGGCGUGGCAGACCACUAAUAGCA .(((.((((..(((...((((((----((((.....((((((...........-..).))))))))))))-))).)))..((((....))))))))))). ( -26.62, z-score = -2.75, R) >droSim1.chrX 3693430 94 + 17042790 CGCUUAUUAAACUCAAGAUAAAC----GCCCGAAAGGCAAAACGAAACGAAAA-AGGAUUUGUGGGCGUU-UAUGGAGGCGUGGCAGACCACUAAUAGCA .(((.((((..(((...((((((----((((....(......)...(((((..-....))))))))))))-))).)))..((((....))))))))))). ( -28.70, z-score = -3.32, R) >droSec1.super_4 4216833 94 - 6179234 CGCUUAUUAAACUCAAGAUAAAC----GCCCGAAAGGCAAAACGAAACGAAAA-AGGAUUUGUGGGCGUU-UAUGGAGGCGUGGCAGACCACUAAUAGCA .(((.((((..(((...((((((----((((....(......)...(((((..-....))))))))))))-))).)))..((((....))))))))))). ( -28.70, z-score = -3.32, R) >droYak2.chrX 18105559 100 - 21770863 CGCUUAUUAAACUCUAGAUAAACAAACGCCCGAAAGAAUAACGAAAAGAAAAACGGGAUUUGUGGGCGCUGUGAGGAGGCGUGGCAGACCACUAAUAGCA .(((.((((..((((......(((..((((((...((((..((..........))..)))).)))))).)))..))))..((((....))))))))))). ( -26.80, z-score = -2.45, R) >droEre2.scaffold_4690 2152241 93 + 18748788 CGCUUAUUAAACGCAAGAUAAAC----GCCCGGGAGAAAAUCGAAAAGAAAAC--GGAUUUGUGGGCGUU-UGAGGAGGCGUGGCAGACCACUAAUAGCA .(((.((((...((....(((((----(((((.(((....(((.........)--)).))).))))))))-)).....))((((....))))))))))). ( -26.60, z-score = -2.20, R) >consensus CGCUUAUUAAACUCAAGAUAAAC____GCCCGAAAGGCAAACGAAAAGAAAAA_AGGAUUUGUGGGCGUU_UAUGGAGGCGUGGCAGACCACUAAUAGCA .(((.((((..................(((((.(((......................))).))))).............((((....))))))))))). (-15.43 = -15.27 + -0.16)


| Location | 4,781,163 – 4,781,257 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 87.45 |
| Shannon entropy | 0.20938 |
| G+C content | 0.45058 |
| Mean single sequence MFE | -23.52 |
| Consensus MFE | -14.47 |
| Energy contribution | -14.51 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.80 |
| Structure conservation index | 0.62 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.13 |
| SVM RNA-class probability | 0.895959 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 4781163 94 - 22422827 UGCUAUUAGUGGUCUGCCACGCCUCCAUA-AACGCCCACAAAUCCU-UUAUUCUUUUCGUUUGCCUUUCGGGC----GUUUAUCUUGAGUUUAAUAAGCG .(((((((((((....))))..(((.(((-(((((((.(((((...-...........)))))......))))----))))))...)))..)))).))). ( -24.64, z-score = -3.43, R) >droSim1.chrX 3693430 94 - 17042790 UGCUAUUAGUGGUCUGCCACGCCUCCAUA-AACGCCCACAAAUCCU-UUUUCGUUUCGUUUUGCCUUUCGGGC----GUUUAUCUUGAGUUUAAUAAGCG .(((((((((((....))))..(((.(((-(((((((.........-......................))))----))))))...)))..)))).))). ( -24.58, z-score = -3.04, R) >droSec1.super_4 4216833 94 + 6179234 UGCUAUUAGUGGUCUGCCACGCCUCCAUA-AACGCCCACAAAUCCU-UUUUCGUUUCGUUUUGCCUUUCGGGC----GUUUAUCUUGAGUUUAAUAAGCG .(((((((((((....))))..(((.(((-(((((((.........-......................))))----))))))...)))..)))).))). ( -24.58, z-score = -3.04, R) >droYak2.chrX 18105559 100 + 21770863 UGCUAUUAGUGGUCUGCCACGCCUCCUCACAGCGCCCACAAAUCCCGUUUUUCUUUUCGUUAUUCUUUCGGGCGUUUGUUUAUCUAGAGUUUAAUAAGCG .(((((((((((....)))).....((((((((((((...(((..((..........))..))).....))))).)))).......)))..)))).))). ( -21.11, z-score = -1.67, R) >droEre2.scaffold_4690 2152241 93 - 18748788 UGCUAUUAGUGGUCUGCCACGCCUCCUCA-AACGCCCACAAAUCC--GUUUUCUUUUCGAUUUUCUCCCGGGC----GUUUAUCUUGCGUUUAAUAAGCG .(((((((((((....))))((......(-(((((((..(((((.--(........).)))))......))))----)))).....))...)))).))). ( -22.70, z-score = -2.80, R) >consensus UGCUAUUAGUGGUCUGCCACGCCUCCAUA_AACGCCCACAAAUCCU_UUUUUCUUUUCGUUUGCCUUUCGGGC____GUUUAUCUUGAGUUUAAUAAGCG .(((((((((((....))))..(((.....(((((((................................))))....)))......)))..)))).))). (-14.47 = -14.51 + 0.04)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:15:52 2011