
| Sequence ID | dm3.chrX |
|---|---|
| Location | 4,732,696 – 4,732,753 |
| Length | 57 |
| Max. P | 0.950104 |

| Location | 4,732,696 – 4,732,753 |
|---|---|
| Length | 57 |
| Sequences | 4 |
| Columns | 57 |
| Reading direction | forward |
| Mean pairwise identity | 95.61 |
| Shannon entropy | 0.07116 |
| G+C content | 0.53070 |
| Mean single sequence MFE | -14.20 |
| Consensus MFE | -13.91 |
| Energy contribution | -13.73 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.98 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.56 |
| SVM RNA-class probability | 0.950104 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 4732696 57 + 22422827 CUUUUUCCGCAUUUUCCGCAGCACUUCAGCCGAGUGAAUGAAUCCUUCACAGCGGAG .....(((((....((.((.........)).))(((((.......))))).))))). ( -14.20, z-score = -1.89, R) >droSim1.chrX 3647855 57 + 17042790 CAUUUUCCGCAUUUUCCGCAGCACUCCAGCCGAGUGAAUGAAUCCUUCACAGCGGAG ..............(((((..(((((.....)))))..(((.....)))..))))). ( -14.20, z-score = -2.13, R) >droYak2.chrX 18056703 57 - 21770863 CAUUUUCCGCAUUUUCCGCAGCACUCCAGCCGAGUGAAUGAAUCCUUCACAGCGGAG ..............(((((..(((((.....)))))..(((.....)))..))))). ( -14.20, z-score = -2.13, R) >droEre2.scaffold_4690 2101168 57 + 18748788 CACUCUCGGCAUUUUCCGCAGCACUCCAGCCGAGUGAAUGAAUCCUUCACAGCGGAG ..............(((((..(((((.....)))))..(((.....)))..))))). ( -14.20, z-score = -1.03, R) >consensus CAUUUUCCGCAUUUUCCGCAGCACUCCAGCCGAGUGAAUGAAUCCUUCACAGCGGAG ..............(((((..(((((.....)))))..(((.....)))..))))). (-13.91 = -13.73 + -0.19)

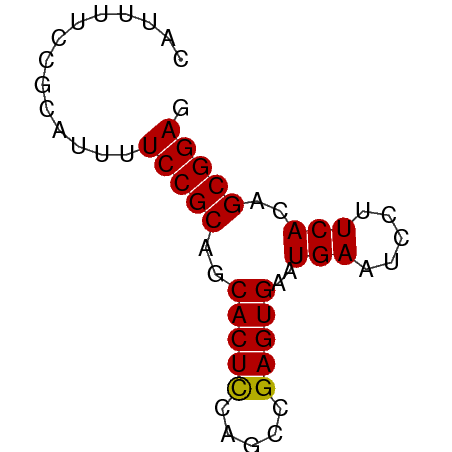
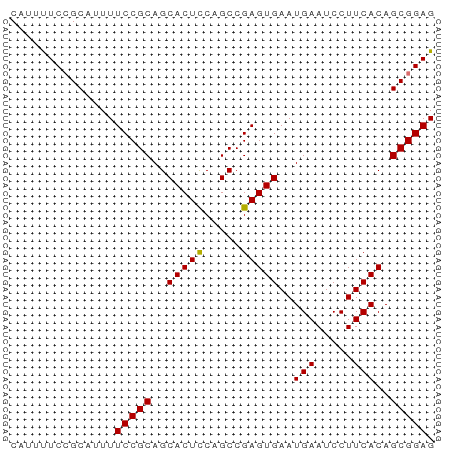
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:15:45 2011