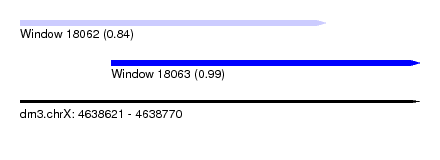
| Sequence ID | dm3.chrX |
|---|---|
| Location | 4,638,621 – 4,638,770 |
| Length | 149 |
| Max. P | 0.992837 |

| Location | 4,638,621 – 4,638,735 |
|---|---|
| Length | 114 |
| Sequences | 11 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.57 |
| Shannon entropy | 0.09250 |
| G+C content | 0.29355 |
| Mean single sequence MFE | -25.39 |
| Consensus MFE | -23.05 |
| Energy contribution | -22.97 |
| Covariance contribution | -0.09 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.91 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.89 |
| SVM RNA-class probability | 0.844667 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 4638621 114 + 22422827 CAUUUUG-CG-----AAAAAUGAUUAAUGCGAUUUGAACUUAAAAUUUACAUUGUGUUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUG ((((((.-..-----.))))))(((((.(((((((.......(((((((.((.((((((((((.((((....)))))))))))))).)).)))))))......)))))))....))))). ( -25.72, z-score = -1.92, R) >droSec1.super_4 4080841 114 - 6179234 CAUUUUA-CG-----AAAAAUGAUUAAUGCGAUUUGAACUUAAAAUUUACAUUGUGGUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUG ((((((.-..-----.))))))(((((.(((((((.......(((((((.((.(((.((((((.((((....)))))))))).))).)).)))))))......)))))))....))))). ( -22.72, z-score = -1.11, R) >droYak2.chrX 17963097 114 - 21770863 CAUUUUG-CG-----AAAAAUGAUUAAUGCGAUUUGAACUUAAAAUUUACAUUGUGUUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUCGAGUUUGUAAAUGAGUUGCUCAUUUAAUG ((((((.-..-----.))))))........(((((((((..............((((((((((.((((....)))))))))))))).)))))))))..((((((((...))))))))... ( -25.46, z-score = -1.59, R) >droEre2.scaffold_4690 2013981 114 + 18748788 CAUUUUG-CG-----AAAAAUGAUUAAUGCGAUUUGAACUUAAAAUUUACAUUGUGUUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUG ((((((.-..-----.))))))(((((.(((((((.......(((((((.((.((((((((((.((((....)))))))))))))).)).)))))))......)))))))....))))). ( -25.72, z-score = -1.92, R) >droAna3.scaffold_12929 3224496 119 + 3277472 -CCUUUGGCGAGCGCAAAAAUGAUUAAUGCGAUUUGAACUUUAAAUUUACAUUGUGUUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUG -.......((((((((...........)))).)))).....((((((((.((.((((((((((.((((....)))))))))))))).)).))))))))((((((((...))))))))... ( -26.70, z-score = -1.34, R) >dp4.chrXL_group1a 2103245 114 - 9151740 CCCUUUG-CG-----AAAAAUGAUUAAUGCGAUUUGAACUUUAAAUUUACAUUGUGUUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUG .......-..-----..((((((.....(((((((......((((((((.((.((((((((((.((((....)))))))))))))).)).)))))))).....))))))))))))).... ( -24.90, z-score = -2.41, R) >droPer1.super_13 639385 114 - 2293547 CCCUUUG-CG-----AAAAAUGAUUAAUGCGAUUUGAACUUUAAAUUUACAUUGUGUUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUG .......-..-----..((((((.....(((((((......((((((((.((.((((((((((.((((....)))))))))))))).)).)))))))).....))))))))))))).... ( -24.90, z-score = -2.41, R) >droWil1.scaffold_181096 7958395 114 - 12416693 -CCUUUUGCG-----AAAAAUGAUUAAUGCGAUUUGAACUUUAAAUUUACAUUGUGUUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUG -.........-----..((((((.....(((((((......((((((((.((.((((((((((.((((....)))))))))))))).)).)))))))).....))))))))))))).... ( -24.90, z-score = -2.13, R) >droVir3.scaffold_12970 7823313 114 - 11907090 CAGUUUG-CG-----AAAAAUGAUUAAUGCGAUUUGAACUUUAAAUUUACAUUGUGUUCAUGAUAUUCAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUG .......-..-----..((((((.....(((((((......((((((((.((.((((((((((.((((....)))))))))))))).)).)))))))).....))))))))))))).... ( -27.30, z-score = -1.96, R) >droMoj3.scaffold_6473 8307438 114 - 16943266 CAGUUUG-CG-----AAAAAUGAUUAAUGCGAUUUGAACUUUAAAUUUACAUUGUGUUCAUGAUAUUCAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUG .......-..-----..((((((.....(((((((......((((((((.((.((((((((((.((((....)))))))))))))).)).)))))))).....))))))))))))).... ( -27.30, z-score = -1.96, R) >droGri2.scaffold_15203 9854841 115 + 11997470 CAGUUUGGCG-----AAAAAUGAUUAAUGCGAUUUGAACUUUAAAUUUACAUUGUGUUCAUGAUAUUCAGUUGAAUUUAUGAGCGUUGUUUGAGUUUGUAAAUGAGUCACUCAUUUAAUG .((((..(((-----....((.....)).))..)..)))).((((((((.((..(((((((((.((((....)))))))))))))..)).))))))))((((((((...))))))))... ( -23.70, z-score = -0.75, R) >consensus CACUUUG_CG_____AAAAAUGAUUAAUGCGAUUUGAACUUUAAAUUUACAUUGUGUUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUG .................((((((.....(((((((.......(((((((.((.((((((((((.((((....)))))))))))))).)).)))))))......))))))))))))).... (-23.05 = -22.97 + -0.09)
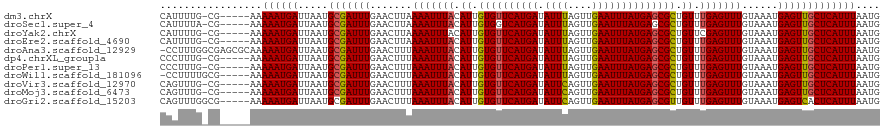
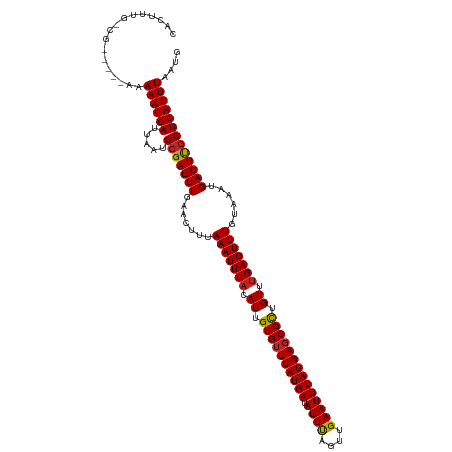
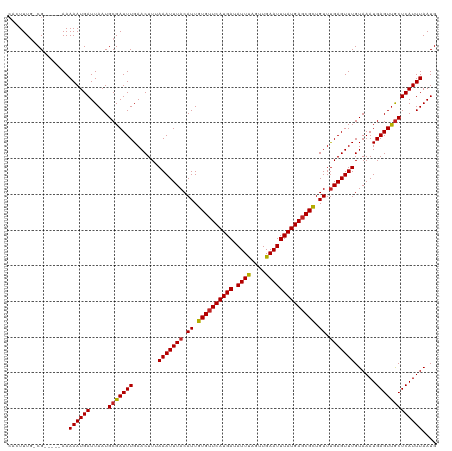
| Location | 4,638,655 – 4,638,770 |
|---|---|
| Length | 115 |
| Sequences | 11 |
| Columns | 125 |
| Reading direction | forward |
| Mean pairwise identity | 81.34 |
| Shannon entropy | 0.38490 |
| G+C content | 0.34503 |
| Mean single sequence MFE | -24.01 |
| Consensus MFE | -20.92 |
| Energy contribution | -20.82 |
| Covariance contribution | -0.10 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.22 |
| Structure conservation index | 0.87 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.57 |
| SVM RNA-class probability | 0.992837 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
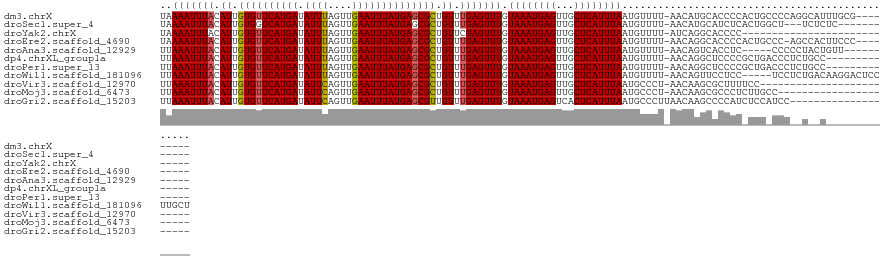
>dm3.chrX 4638655 115 + 22422827 UAAAAUUUACAUUGUGUUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUGUUUU-AACAUGCACCCCACUGCCCCAGGCAUUUGCG--------- ..(((((((.((.((((((((((.((((....)))))))))))))).)).))))))).((((((((...))))))))((((...-.))))(((......((((...))))..))).--------- ( -26.20, z-score = -1.71, R) >droSec1.super_4 4080875 109 - 6179234 UAAAAUUUACAUUGUGGUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUGUUUU-AACAUGCAUCUCACUGGCU---UCUCUC------------ ..(((((((.((.(((.((((((.((((....)))))))))).))).)).)))))))((...((((.(((....((((....))-))...))).))))...)).---......------------ ( -21.70, z-score = -0.55, R) >droYak2.chrX 17963131 96 - 21770863 UAAAAUUUACAUUGUGUUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUCGAGUUUGUAAAUGAGUUGCUCAUUUAAUGUUUU-AUCAGGCACCCC---------------------------- .............((((((((((.((((....))))))))))))))(((..((.....((((((((...)))))))).......-.))..)))....---------------------------- ( -20.52, z-score = -1.77, R) >droEre2.scaffold_4690 2014015 114 + 18748788 UAAAAUUUACAUUGUGUUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUGUUUU-AACAGGCACCCCACUGCCC-AGCCACUUCCC--------- ..(((((((.((.((((((((((.((((....)))))))))))))).)).))))))).((((((((...)))))))).......-....((((......)))).-...........--------- ( -25.60, z-score = -2.17, R) >droAna3.scaffold_12929 3224535 108 + 3277472 UUAAAUUUACAUUGUGUUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUGUUUU-AACAGUCACCUC-----CCCCCUACUGUU----------- .((((((((.((.((((((((((.((((....)))))))))))))).)).))))))))((((((((...)))))))).......-((((((......-----......))))))----------- ( -24.30, z-score = -3.47, R) >dp4.chrXL_group1a 2103279 110 - 9151740 UUAAAUUUACAUUGUGUUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUGUUUU-AACAGGCUCCCCGCUGACCCUCUGCC-------------- .((((((((.((.((((((((((.((((....)))))))))))))).)).))))))))((((((((...)))))))).......-...(((.((......)).))).....-------------- ( -23.80, z-score = -1.97, R) >droPer1.super_13 639419 110 - 2293547 UUAAAUUUACAUUGUGUUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUGUUUU-AACAGGCUCCCCGCUGACCCUCUGCC-------------- .((((((((.((.((((((((((.((((....)))))))))))))).)).))))))))((((((((...)))))))).......-...(((.((......)).))).....-------------- ( -23.80, z-score = -1.97, R) >droWil1.scaffold_181096 7958429 119 - 12416693 UUAAAUUUACAUUGUGUUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUGUUUU-AACAGUUCCUCC-----UCCUCUGACAAGGACUCCUUGCU .((((((((.((.((((((((((.((((....)))))))))))))).)).))))))))((((((((...)))))))).......-..(((.......-----....))).(((((...))))).. ( -26.10, z-score = -1.98, R) >droVir3.scaffold_12970 7823347 99 - 11907090 UUAAAUUUACAUUGUGUUCAUGAUAUUCAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUGCCCU-AACAAGCGCUUUUCC------------------------- .((((((((.((.((((((((((.((((....)))))))))))))).)).))))))))((((((((...))))))))..((.((-....)).))......------------------------- ( -25.30, z-score = -3.20, R) >droMoj3.scaffold_6473 8307472 102 - 16943266 UUAAAUUUACAUUGUGUUCAUGAUAUUCAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUGCCCU-AACAAGCGCCCUCUUGCC---------------------- .((((((((.((.((((((((((.((((....)))))))))))))).)).))))))))((((((((...)))))))).......-..((((......))))..---------------------- ( -25.30, z-score = -2.98, R) >droGri2.scaffold_15203 9854876 105 + 11997470 UUAAAUUUACAUUGUGUUCAUGAUAUUCAGUUGAAUUUAUGAGCGUUGUUUGAGUUUGUAAAUGAGUCACUCAUUUAAUGCCCUUAACAAGCCCCAUCUCCAUCC-------------------- .((((((((.((..(((((((((.((((....)))))))))))))..)).))))))))((((((((...))))))))............................-------------------- ( -21.50, z-score = -2.67, R) >consensus UUAAAUUUACAUUGUGUUCAUGAUAUUUAGUUGAAUUUAUGAGCGCUGUUUGAGUUUGUAAAUGAGUUGCUCAUUUAAUGUUUU_AACAGGCACCCC_CUG_CCC__U__C______________ ..(((((((.((.((((((((((.((((....)))))))))))))).)).))))))).((((((((...))))))))................................................ (-20.92 = -20.82 + -0.10)

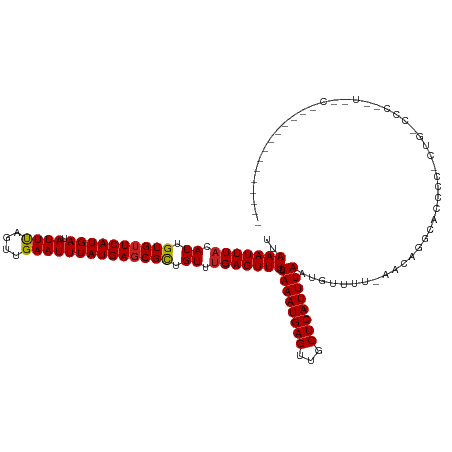
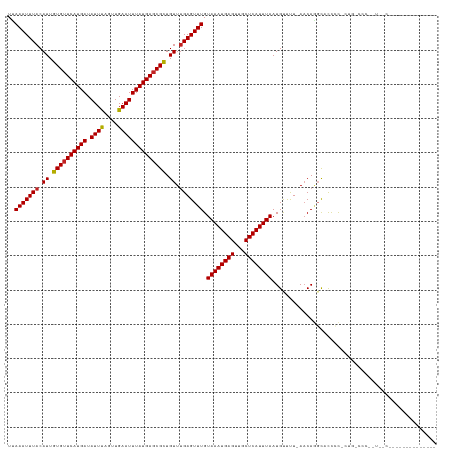
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:15:25 2011