
| Sequence ID | dm3.chrX |
|---|---|
| Location | 4,470,308 – 4,470,404 |
| Length | 96 |
| Max. P | 0.873692 |

| Location | 4,470,308 – 4,470,404 |
|---|---|
| Length | 96 |
| Sequences | 8 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 54.98 |
| Shannon entropy | 0.83383 |
| G+C content | 0.50424 |
| Mean single sequence MFE | -25.84 |
| Consensus MFE | -8.60 |
| Energy contribution | -8.24 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.73 |
| Mean z-score | -1.35 |
| Structure conservation index | 0.33 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.873692 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 4470308 96 + 22422827 CAGCCCGCAGACCAAUCCAAUCCAAUACAAUACAAACCGAU--GGCUUUUAUGAGCACAUGGGCUUCCAUGGUCCCAAAUAGUUGGGCCUCCGGGCUG--- (((((((.............................((.((--(((((....)))).))).)).......((.(((((....)))))))..)))))))--- ( -31.00, z-score = -2.01, R) >droSim1.chrX 3472843 79 + 17042790 ------------CAAUCCA-----AUACAAUACAAUCCGAU--GGCUUUUAUGAGCACAUGGGCUUCCAUGGGCCCAAAUAGUUGGGCCUCCGGGCUG--- ------------.......-----........((..(((..--.((((....)))).(((((....)))))(((((((....)))))))..)))..))--- ( -27.40, z-score = -1.92, R) >droSec1.super_4 3921359 79 - 6179234 ------------CAAUCCA-----AUACAAUACAAUCCGAU--GGCUUUUAUGAGCACAUGGGCUUCCAUGGGCCCAAAUAGUUGGGCCUCCGGGCUG--- ------------.......-----........((..(((..--.((((....)))).(((((....)))))(((((((....)))))))..)))..))--- ( -27.40, z-score = -1.92, R) >droYak2.chrX 17803792 91 - 21770863 CAGCCCGCAGACCAAUCCA-----AUACAAUACAAUCCGAU--GGCUUUUAUGAGCACAUGGGCUUCCAUGGGCCCAAAUAGUUGGGCCUCCGGGCUG--- (((((((............-----............((.((--(((((....)))).))).)).......((((((((....)))))))).)))))))--- ( -35.10, z-score = -2.77, R) >droEre2.scaffold_4690 1849623 93 + 18748788 CAGUCCGCAGACCAAUCCA-----AUACAAUACAAUCCGAUCCGGCUUUUAUGAGCACAUGGGCUUCCAUGGGCCCAAAUAGUUGGGCCUCCGGGCUG--- (((((((............-----....................((((....)))).(((((....)))))(((((((....)))))))..)))))))--- ( -31.50, z-score = -1.85, R) >dp4.chrXL_group1a 358560 86 - 9151740 ------AUGGCCAUGCCCAAG--ACUCCACUCCACUCCCAU---CCCCCUGCUGUUGUCUGG-CUUUUACGAGCCCGAAUAGUUGGGCCUCUGUGCUG--- ------..(((((.(((((((--((..(((...........---......).))..))))((-(((....)))))........)))))...)).))).--- ( -20.03, z-score = -0.24, R) >droVir3.scaffold_13042 2911275 84 + 5191987 -----------------CAUUGUCCAUUGUCGAUUACCCAUAGACCUGUGUCUGGUCUCUGAUUUGCUCUUAGAAUUUUAAACAGGCUCUUCAGAUGAUGA -----------------.......(((..((((....((...((((.......))))(((((.......)))))..........))....)).))..))). ( -15.80, z-score = 0.21, R) >droMoj3.scaffold_6328 3731179 89 - 4453435 ----------CACUGCCCAUUGCGCAUUGCCCAUAGACCAUAGACCUUUGUCUGGUCUCUGAUUUGCUCUCAGAAUUUUAAAUAGGCUCUUCAGAUGAU-- ----------.......((((.......(((....(((((..(((....))))))))(((((.......)))))..........)))......))))..-- ( -18.52, z-score = -0.31, R) >consensus ___________CCAAUCCA_____AUACAAUACAAUCCGAU__GGCUUUUAUGAGCACAUGGGCUUCCAUGGGCCCAAAUAGUUGGGCCUCCGGGCUG___ ...........................................................(((((((....)))))))........((((....)))).... ( -8.60 = -8.24 + -0.36)


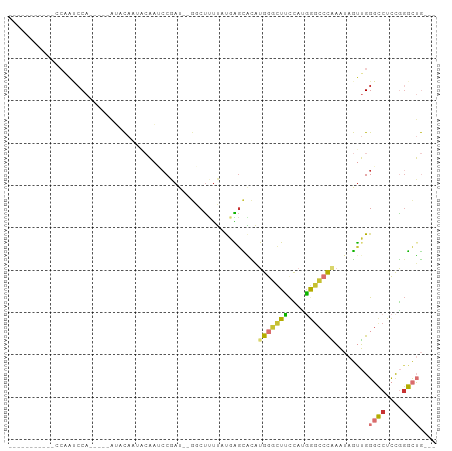
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:15:10 2011