
| Sequence ID | dm3.chrX |
|---|---|
| Location | 4,466,716 – 4,466,809 |
| Length | 93 |
| Max. P | 0.989370 |

| Location | 4,466,716 – 4,466,809 |
|---|---|
| Length | 93 |
| Sequences | 8 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 88.42 |
| Shannon entropy | 0.23899 |
| G+C content | 0.44065 |
| Mean single sequence MFE | -27.11 |
| Consensus MFE | -21.26 |
| Energy contribution | -21.73 |
| Covariance contribution | 0.47 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.86 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.36 |
| SVM RNA-class probability | 0.989370 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 4466716 93 + 22422827 AUCCAUCCAUCUGUUUGGUCGGCUGACUCUUGGCUGCGUUGCAUGUUUGAUUUCU-GCUAAUGCAAUGCGGCAUCUUGUUGCUGUUGUAAUUAA ..(((..(....)..))).((((.(((.....((((((((((((....(....).-....)))))))))))).....))))))).......... ( -27.90, z-score = -2.64, R) >droSim1.chrX 3469346 93 + 17042790 UCCAAUCCAUCUGUUUGGUCGGCUGACUCUUGGCUGCGUUGCAUGUUUGAUUUCU-GCUAAUGCAAUGCGGCAUCUUGUUGCUGUUGUAAUUAA ......(((......))).((((.(((.....((((((((((((....(....).-....)))))))))))).....))))))).......... ( -27.20, z-score = -2.43, R) >droSec1.super_4 3917701 93 - 6179234 UCAAAUCCAUCUGUUUGGUCGGCUGACUCUUGGCUGCGUUGCAUGUUUGAUUUCU-GCUAAUGCAAUGCGGCAUCUUGUUGCUGUUGUAAUUAA ......(((......))).((((.(((.....((((((((((((....(....).-....)))))))))))).....))))))).......... ( -27.20, z-score = -2.54, R) >droYak2.chrX 17800300 89 - 21770863 ----AUCCAUCUGUUUGGUCGGCUGACUCUUGGCUGCGUUGCAUGUUUGAUUUCU-GCUAAUGCAAUGCGGCAUCUUGUUGCUGUUGUAAUUAA ----..(((......))).((((.(((.....((((((((((((....(....).-....)))))))))))).....))))))).......... ( -27.20, z-score = -2.84, R) >droEre2.scaffold_4690 1846056 89 + 18748788 ----AUCCAUCUGUUUGGUCGGCUGACUCUUGGCUGCGUUGCAUGUUUGAUUUCU-GCUAAUGCAAUGCGGCAUCUUGUUGCUGUUGUAAUUAA ----..(((......))).((((.(((.....((((((((((((....(....).-....)))))))))))).....))))))).......... ( -27.20, z-score = -2.84, R) >droAna3.scaffold_13335 1909628 78 + 3335858 ----GUCCAUCUGCCUGCUUGGCUG---CUUGGCUGCGUUGCAUGUUUGAUUUUUUGUUAAUGCAAUGCGGCAUCUUGUAAUUAA--------- ----........(((.....)))((---(...((((((((((((...(((....)))...)))))))))))).....))).....--------- ( -24.30, z-score = -2.41, R) >dp4.chrXL_group1a 8797634 91 + 9151740 -UCCAUCCAUCUGUCUGCUCGGCUGCCUCUUG-CUGCGUUGCAUGUUUGAUUUCU-GUUAAUGCAAUGCGGCAUCUUGUUGCUGUUGUAAUUAA -..............(((.((((.((....((-(((((((((((...((((....-)))))))))))))))))....)).))))..)))..... ( -28.20, z-score = -3.99, R) >droPer1.super_15 1191212 91 + 2181545 -UCCAUCCAUCUGUCUGCUCGGCUGGCUCUUG-CUGCGUUGCAUGUUUGAUUUCU-GUUAAUGCAAUGCGGCAUCUUGUUGCUGUUGUAAUUAA -..............(((.((((.(((...((-(((((((((((...((((....-)))))))))))))))))....)))))))..)))..... ( -27.70, z-score = -3.20, R) >consensus __C_AUCCAUCUGUUUGGUCGGCUGACUCUUGGCUGCGUUGCAUGUUUGAUUUCU_GCUAAUGCAAUGCGGCAUCUUGUUGCUGUUGUAAUUAA ...................((((.(((.....((((((((((((................)))))))))))).....))))))).......... (-21.26 = -21.73 + 0.47)


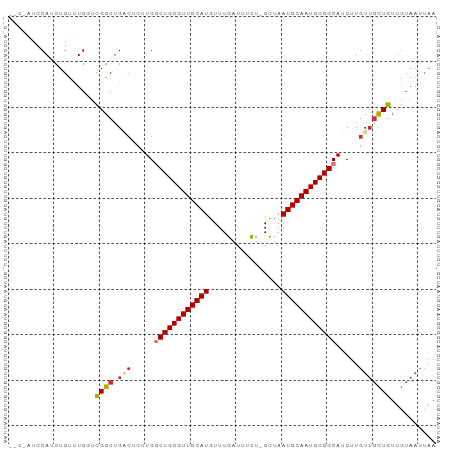
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:15:08 2011