
| Sequence ID | dm3.chrX |
|---|---|
| Location | 4,310,646 – 4,310,756 |
| Length | 110 |
| Max. P | 0.515939 |

| Location | 4,310,646 – 4,310,756 |
|---|---|
| Length | 110 |
| Sequences | 8 |
| Columns | 128 |
| Reading direction | reverse |
| Mean pairwise identity | 76.73 |
| Shannon entropy | 0.43362 |
| G+C content | 0.33085 |
| Mean single sequence MFE | -18.07 |
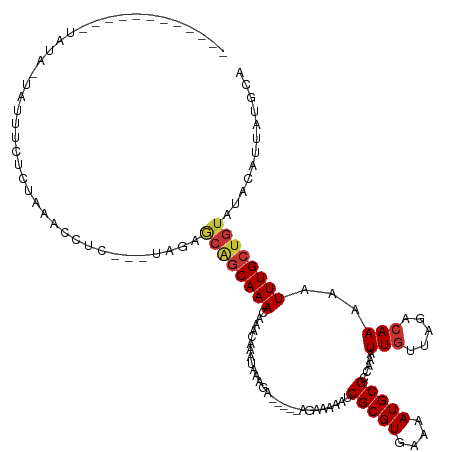
| Consensus MFE | -13.73 |
| Energy contribution | -14.04 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.12 |
| Mean z-score | -0.70 |
| Structure conservation index | 0.76 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.515939 |
| Prediction | RNA |
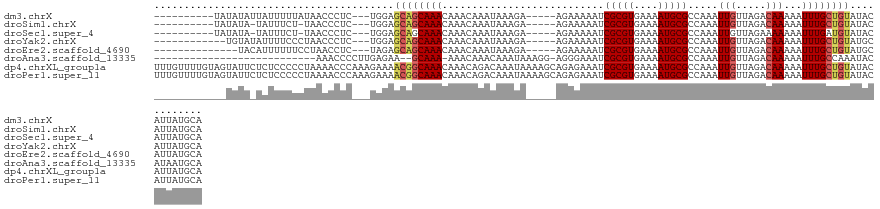
Download alignment: ClustalW | MAF
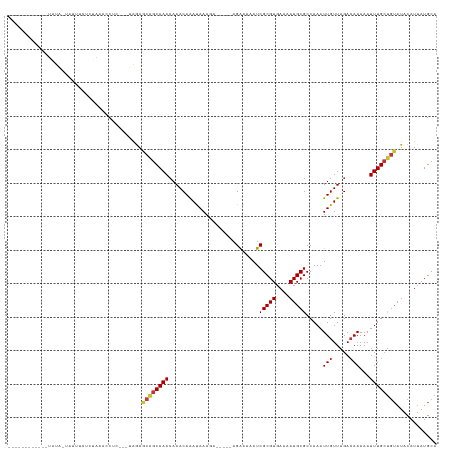
>dm3.chrX 4310646 110 - 22422827 ----------UAUAUAUUAUUUUUAUAACCCUC---UGGAGCAGCAAACAAACAAAUAAAGA-----AGAAAAAUCGCGUGAAAAUGCGCCAAAUUGUUAGACAAAAAUUUGCUGUAUACAUUAUGCA ----------.............(((((((...---.)).((((((((..............-----........(((((....))))).....(((.....)))...)))))))).....))))).. ( -16.80, z-score = -0.46, R) >droSim1.chrX 3319906 108 - 17042790 ----------UAUAUA-UAUUUCU-UAACCCUC---UGGAGCAGCAAACAAACAAAUAAAGA-----AGAAAAAUCGCGUGAAAAUGCGCCAAAUUGUUAGACAAAAAUUUGCUGUAUACAUUAUGCA ----------......-.......-...((...---.)).((((((((..............-----........(((((....))))).....(((.....)))...))))))))............ ( -16.60, z-score = -0.55, R) >droSec1.super_4 3763876 108 + 6179234 ----------UAUAUA-UAUUUCU-UAACCCUC---UGGAGCAGCAAACAAACAAAUAAAGA-----AGAAAAAUCGCGUGAAAAUGCGCCAAAUUGUUAGAAAAAAAUUUGAUGUAUACAUUAUGCA ----------......-..(((((-(.......---((..............)).......)-----)))))....((((((..(((((.((((((..........)))))).)))))...)))))). ( -14.78, z-score = -0.19, R) >droYak2.chrX 17643002 108 + 21770863 ------------UGUAUAUUUUCCCUAACCCUC---UGGAGCAGCAAACAAACAAAUAAAGA-----AGAAAAAUCGCGUGAAAAUGCGCCAAAUUGUUAGACAAAAAUUUGCUGUAUGCAUUAUGCA ------------.(((((...(((.........---.)))((((((((..............-----........(((((....))))).....(((.....)))...))))))))......))))). ( -20.30, z-score = -1.15, R) >droEre2.scaffold_4690 1687044 106 - 18748788 --------------UACAUUUUUUCCUAACCUC---UAGAGCAGCAAACAAACAAAUAAAGA-----AGAAAAAUCGCGUGAAAAUGCGCCAAAUUGUUAGACAAAAAUUUGCUGUAUGCAUUAUGCA --------------...................---....((((((((..............-----........(((((....))))).....(((.....)))...)))))))).(((.....))) ( -18.00, z-score = -0.73, R) >droAna3.scaffold_13335 3183162 97 + 3335858 ---------------------------AAACCCCUUGAGAA--GCAAA-AAACAAACAAAUAAAGG-AGGGAAAUCGCGUGAAAAUGCGCCAAAUUGUUAGACAAAAAUUUGCCAAAUACAUAAUGCA ---------------------------...((((((.....--.....-.............))))-.)).....(((((....))))).((((((..........))))))................ ( -12.80, z-score = 0.12, R) >dp4.chrXL_group1a 5406369 128 + 9151740 UUUGUUUUGUAGUAUUCUCUCCCCCUAAAACCCAAAGAAAACGGCAAACAAACAGACAAAUAAAAGCAGAGAAAUCGCGUGAAAAUGCGCCAAAUUGUUAGACAAAAAUUUGCUGUAUACAUUAUGCA .((((((.((.((.((((.................)))).)).))))))))((((.(((((....((.((....))((((....))))))....(((.....)))..)))))))))............ ( -22.63, z-score = -1.31, R) >droPer1.super_11 2540917 128 + 2846995 UUUGUUUUGUAGUAUUCUCUCCCCCUAAAACCCAAAGAAAACGGCAAACAAACAGACAAAUAAAAGCAGAGAAAUCGCGUGAAAAUGCGCCAAAUUGUUAGACAAAAAUUUGCUGUAUACAUUAUGCA .((((((.((.((.((((.................)))).)).))))))))((((.(((((....((.((....))((((....))))))....(((.....)))..)))))))))............ ( -22.63, z-score = -1.31, R) >consensus ____________UAUA_UAUUUCUCUAAACCUC___UAGAGCAGCAAACAAACAAAUAAAGA_____AGAAAAAUCGCGUGAAAAUGCGCCAAAUUGUUAGACAAAAAUUUGCUGUAUACAUUAUGCA ........................................((((((((...........................(((((....))))).....(((.....)))...))))))))............ (-13.73 = -14.04 + 0.31)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:14:40 2011