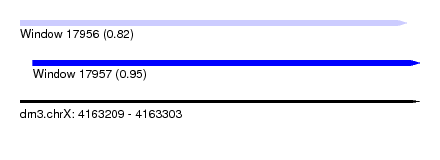

| Sequence ID | dm3.chrX |
|---|---|
| Location | 4,163,209 – 4,163,303 |
| Length | 94 |
| Max. P | 0.946601 |

| Location | 4,163,209 – 4,163,300 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 82.78 |
| Shannon entropy | 0.33255 |
| G+C content | 0.39825 |
| Mean single sequence MFE | -15.47 |
| Consensus MFE | -9.33 |
| Energy contribution | -9.78 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.38 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.820927 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
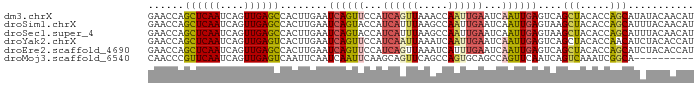
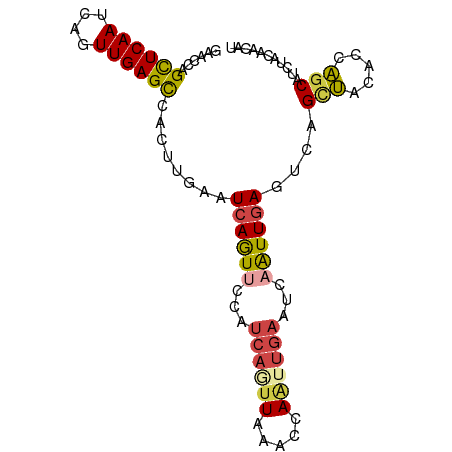
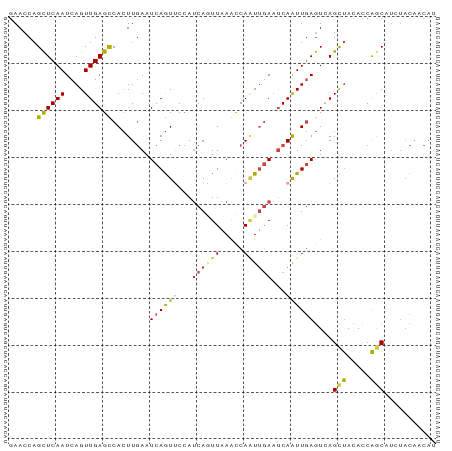
>dm3.chrX 4163209 91 + 22422827 GAACCAGCUCAAUCAGUUGAGCCACUUGAAUCAGUUCCAUCAGUUAAACCAAUUGAAUCAAUUGAGUCAGCUACACCAGCAUAUACAACAU ......((((((....))))))....(((.((((((...((((((.....))))))...)))))).)))(((.....)))........... ( -18.40, z-score = -3.01, R) >droSim1.chrX 3190426 91 + 17042790 GAACCAGCUCAAUCAGUUGAGCCACUUGAAUCAGUACCAUCAUUUAAGCCAAUUGAAUCAAUUGAGUAAGCUACACCAGCAUUUACAACAU .....((((...((((((((..(((((((((.(......).))))))).....))..))))))))...))))................... ( -16.00, z-score = -2.16, R) >droSec1.super_4 3610372 91 - 6179234 GAACCAGCUCAAUCAGUUGAGCCACUUGAAUCAGUACCAUCAUUUAAGCCAAUUGAAUCAAUUGAGUAAGCUACACCAGCAUUUACAACAU .....((((...((((((((..(((((((((.(......).))))))).....))..))))))))...))))................... ( -16.00, z-score = -2.16, R) >droYak2.chrX 17496644 91 - 21770863 GAACCAGCUCAAUCAGUUGAGUCACUUGAAUCAGUUCCAUCAAUUAAAUCAAUUGAAUCAAUUGAGUCAGCUACACCAACAUCUACACCAU .....((((...((((((((.(((.((((.(.((((.....)))).).)))).))).))))))))...))))................... ( -18.20, z-score = -3.11, R) >droEre2.scaffold_4690 1538492 91 + 18748788 GAACCAGCUCAAUCAGUUGAGCCACUUGAAUCAGUUCCAUCAGUUAAAUCAUUUGAAUCAAUUGAGUCAGCUACACCAGCAUCUACACCAU ......((((((....))))))....(((.((((((...((((.........))))...)))))).)))(((.....)))........... ( -16.10, z-score = -1.71, R) >droMoj3.scaffold_6540 16821850 81 - 34148556 CAACCCGUUCAAUCAGUUGAGUCAAUUCAAUCAAUUCAAGCAGUUCAGCCAGUGCAGCCAGUUCAAUCAGUCAAAUCGGCA---------- ..........(((..((((((....))))))..)))...(((.(......).))).(((.(((..........))).))).---------- ( -8.10, z-score = -0.52, R) >consensus GAACCAGCUCAAUCAGUUGAGCCACUUGAAUCAGUUCCAUCAGUUAAACCAAUUGAAUCAAUUGAGUCAGCUACACCAGCAUCUACAACAU ......((((((....))))))........((((((...((((((.....))))))...))))))....(((.....)))........... ( -9.33 = -9.78 + 0.45)
| Location | 4,163,212 – 4,163,303 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 81.61 |
| Shannon entropy | 0.35573 |
| G+C content | 0.40263 |
| Mean single sequence MFE | -15.47 |
| Consensus MFE | -9.33 |
| Energy contribution | -9.78 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.38 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.52 |
| SVM RNA-class probability | 0.946601 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
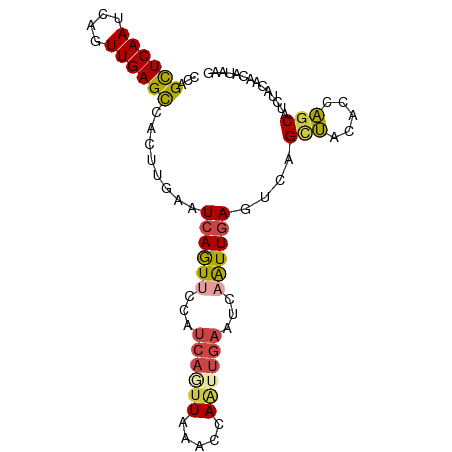
>dm3.chrX 4163212 91 + 22422827 CCAGCUCAAUCAGUUGAGCCACUUGAAUCAGUUCCAUCAGUUAAACCAAUUGAAUCAAUUGAGUCAGCUACACCAGCAUAUACAACAUAAG ...((((((....))))))....(((.((((((...((((((.....))))))...)))))).)))(((.....))).............. ( -18.40, z-score = -3.34, R) >droSim1.chrX 3190429 91 + 17042790 CCAGCUCAAUCAGUUGAGCCACUUGAAUCAGUACCAUCAUUUAAGCCAAUUGAAUCAAUUGAGUAAGCUACACCAGCAUUUACAACAUAAG ..((((...((((((((..(((((((((.(......).))))))).....))..))))))))...))))...................... ( -16.00, z-score = -2.34, R) >droSec1.super_4 3610375 91 - 6179234 CCAGCUCAAUCAGUUGAGCCACUUGAAUCAGUACCAUCAUUUAAGCCAAUUGAAUCAAUUGAGUAAGCUACACCAGCAUUUACAACAUAAG ..((((...((((((((..(((((((((.(......).))))))).....))..))))))))...))))...................... ( -16.00, z-score = -2.34, R) >droYak2.chrX 17496647 91 - 21770863 CCAGCUCAAUCAGUUGAGUCACUUGAAUCAGUUCCAUCAAUUAAAUCAAUUGAAUCAAUUGAGUCAGCUACACCAACAUCUACACCAUCAG ..((((...((((((((.(((.((((.(.((((.....)))).).)))).))).))))))))...))))...................... ( -18.20, z-score = -3.52, R) >droEre2.scaffold_4690 1538495 91 + 18748788 CCAGCUCAAUCAGUUGAGCCACUUGAAUCAGUUCCAUCAGUUAAAUCAUUUGAAUCAAUUGAGUCAGCUACACCAGCAUCUACACCAUCAG ...((((((....))))))....(((.((((((...((((.........))))...)))))).)))(((.....))).............. ( -16.10, z-score = -2.07, R) >droMoj3.scaffold_6540 16821853 78 - 34148556 CCCGUUCAAUCAGUUGAGUCAAUUCAAUCAAUUCAAGCAGUUCAGCCAGUGCAGCCAGUUCAAUCAGUCAAAUCGGCA------------- .......(((..((((((....))))))..)))...(((.(......).))).(((.(((..........))).))).------------- ( -8.10, z-score = -0.68, R) >consensus CCAGCUCAAUCAGUUGAGCCACUUGAAUCAGUUCCAUCAGUUAAACCAAUUGAAUCAAUUGAGUCAGCUACACCAGCAUCUACAACAUAAG ...((((((....))))))........((((((...((((((.....))))))...))))))....(((.....))).............. ( -9.33 = -9.78 + 0.45)

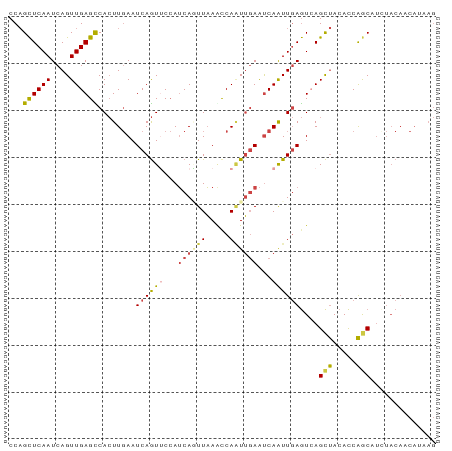
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:13:58 2011