| Sequence ID | dm3.chrX |
|---|---|
| Location | 4,042,211 – 4,042,268 |
| Length | 57 |
| Max. P | 0.797395 |

| Location | 4,042,211 – 4,042,268 |
|---|---|
| Length | 57 |
| Sequences | 5 |
| Columns | 59 |
| Reading direction | reverse |
| Mean pairwise identity | 79.24 |
| Shannon entropy | 0.36883 |
| G+C content | 0.51210 |
| Mean single sequence MFE | -17.10 |
| Consensus MFE | -10.84 |
| Energy contribution | -10.96 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.41 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.797395 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 4042211 57 - 22422827 CUAUCUACUUGCGCCGGAUGCACAAACACAUUUGGUGGUACGGUGUAA--CUGGCCAAG .......((((.(((((.(((((...(((.....))).....))))).--))))))))) ( -21.00, z-score = -3.12, R) >droSim1.chrX 3015626 57 - 17042790 CUAUCUACUUGCGCCGGAUGCACAAACACAUUUGGUGGAAGGGUGUAG--CUGGCCAAG .......((((.(((((.(((((...(((.....))).....))))).--))))))))) ( -21.00, z-score = -2.62, R) >droYak2.chrX 17383708 57 + 21770863 CUAUCUACUUGCGUCGGAUGCACAAACACAUUUGGUGGAAGGGUGUAU--CUGGUCAAG .......((((.(((((((((((...(((.....))).....))))))--))))))))) ( -17.30, z-score = -2.55, R) >droEre2.scaffold_4690 1419141 57 - 18748788 CUAUCUACUUGCGUCGGAUGCACAAACACAUUUGGUGGAAGGGUGUAG--CUGGCCGAG .......((((.(((((.(((((...(((.....))).....))))).--))))))))) ( -18.00, z-score = -1.80, R) >droAna3.scaffold_13117 216723 59 + 5790199 CUAUCUACCUUCACCUUGUACUUGCACCCAUCCACUGGCAGGAUCUGGCUCUGGCAAAG ...............((((....((....((((.......))))...))....)))).. ( -8.20, z-score = 1.20, R) >consensus CUAUCUACUUGCGCCGGAUGCACAAACACAUUUGGUGGAAGGGUGUAG__CUGGCCAAG .......((((.(((((.(((((...(((.....))).....)))))...))))))))) (-10.84 = -10.96 + 0.12)

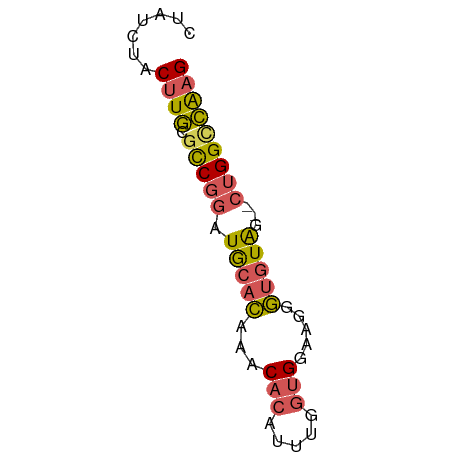

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:13:30 2011