
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 9,975,542 – 9,975,649 |
| Length | 107 |
| Max. P | 0.989428 |

| Location | 9,975,542 – 9,975,649 |
|---|---|
| Length | 107 |
| Sequences | 3 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 55.14 |
| Shannon entropy | 0.64895 |
| G+C content | 0.40831 |
| Mean single sequence MFE | -28.00 |
| Consensus MFE | -10.75 |
| Energy contribution | -10.20 |
| Covariance contribution | -0.55 |
| Combinations/Pair | 1.27 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.37 |
| SVM RNA-class probability | 0.989428 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 9975542 107 + 23011544 AAAUAAUUUAGCGGACUUUGACAGGGUGCCUGAUAUCGUAAAAUCGAUCAGGGAAUUCUCUGGAAAUCCAGCAGAAUUCGGGUCUUGGAAAAAGAGCGUUGAUAGAA .......((((((..((((..(((((..(((((..(((......))))))))(((((((((((....)))).)))))))...)))))...))))..))))))..... ( -34.70, z-score = -3.40, R) >droAna3.scaffold_13034 1551495 94 - 2223656 -------------GACUUAGAUCGGGUACCAGAUAUAGUGAAGUGCAUGAAGGAAUUUUCGGGAAAGCCCACAGAAUUUAGUUCGUGGACAAAAACGGUAGAUCGAG -------------......((((((...))...(((.((...((.((((((.(((((((.(((....)))..)))))))..)))))).))....)).)))))))... ( -21.30, z-score = -1.18, R) >droWil1.scaffold_180943 117849 98 + 149748 AGAUAAUUGGCAUGAGGUAGAUAAAGUUCCAGACUUAGUGAAAGGUCUAAAGGAAUUCUCUGGUAAUCCAGCAGAAUUUACAACUUGGUUCGCUUCAA--------- ............(((((..(((.(((((..((((((......))))))....(((((((((((....)))).)))))))..))))).)))..))))).--------- ( -28.00, z-score = -2.56, R) >consensus A_AUAAUU_____GACUUAGAUAGGGUACCAGAUAUAGUGAAAUGCAUAAAGGAAUUCUCUGGAAAUCCAGCAGAAUUUAGAUCUUGGAAAAAAACAGU_GAU_GA_ ....................................................(((((((((((....)))).)))))))............................ (-10.75 = -10.20 + -0.55)


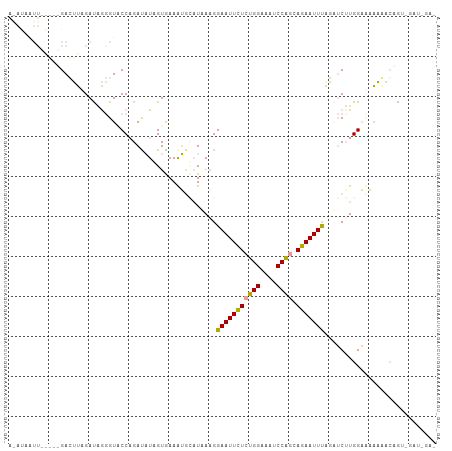
| Location | 9,975,542 – 9,975,649 |
|---|---|
| Length | 107 |
| Sequences | 3 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 55.14 |
| Shannon entropy | 0.64895 |
| G+C content | 0.40831 |
| Mean single sequence MFE | -18.70 |
| Consensus MFE | -7.70 |
| Energy contribution | -7.37 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.24 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.712776 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 9975542 107 - 23011544 UUCUAUCAACGCUCUUUUUCCAAGACCCGAAUUCUGCUGGAUUUCCAGAGAAUUCCCUGAUCGAUUUUACGAUAUCAGGCACCCUGUCAAAGUCCGCUAAAUUAUUU .........((..((((......(((..(((((((.((((....)))))))))))((((((((......)))..)))))......)))))))..))........... ( -23.70, z-score = -2.33, R) >droAna3.scaffold_13034 1551495 94 + 2223656 CUCGAUCUACCGUUUUUGUCCACGAACUAAAUUCUGUGGGCUUUCCCGAAAAUUCCUUCAUGCACUUCACUAUAUCUGGUACCCGAUCUAAGUC------------- ...((((((((......(((((((((.....))).))))))......(((......)))..................))))...))))......------------- ( -17.20, z-score = -1.53, R) >droWil1.scaffold_180943 117849 98 - 149748 ---------UUGAAGCGAACCAAGUUGUAAAUUCUGCUGGAUUACCAGAGAAUUCCUUUAGACCUUUCACUAAGUCUGGAACUUUAUCUACCUCAUGCCAAUUAUCU ---------.(((((..............((((((.((((....))))))))))..(((((((..........))))))).)))))..................... ( -15.20, z-score = 0.13, R) >consensus _UC_AUC_ACCGUAUUUAUCCAAGAACUAAAUUCUGCUGGAUUUCCAGAGAAUUCCUUCAUACAUUUCACUAUAUCUGGAACCCUAUCUAAGUC_____AAUUAU_U .............................((((((.((((....))))))))))..................................................... ( -7.70 = -7.37 + -0.33)

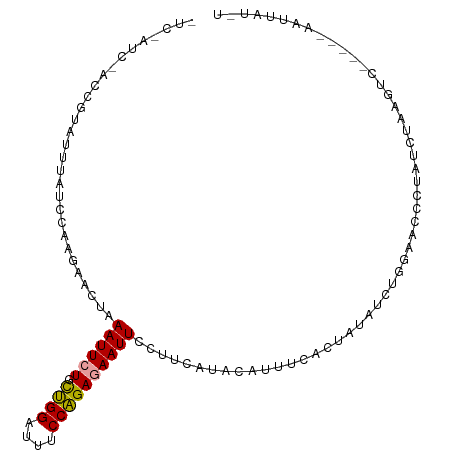
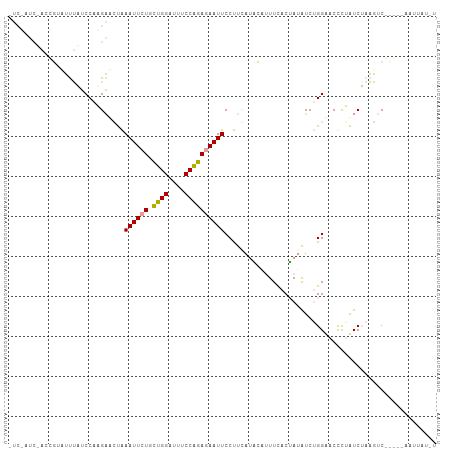
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:29:24 2011