
| Sequence ID | dm3.chrX |
|---|---|
| Location | 3,878,957 – 3,879,052 |
| Length | 95 |
| Max. P | 0.584829 |

| Location | 3,878,957 – 3,879,052 |
|---|---|
| Length | 95 |
| Sequences | 9 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 79.56 |
| Shannon entropy | 0.41890 |
| G+C content | 0.51402 |
| Mean single sequence MFE | -32.94 |
| Consensus MFE | -17.01 |
| Energy contribution | -17.79 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.584829 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 3878957 95 - 22422827 ---GUGCGGCAGC-GGCGGCGACAUCGGC-UAAAGGCAAGGCGCUCAUUAGGAUAAUUGCAGGCAAUUAACUAAAUUGCCGCCGCAUUUGUCGCU---CCUCG ---..((((((((-((((((((.....((-(........))).....((((..((((((....)))))).)))).))))))))))...)))))).---..... ( -37.70, z-score = -2.09, R) >droSec1.super_4 3340123 94 + 6179234 ----GGCGGUAGC-GGCGGCGACAUCGGC-UAAAGGCAAGGCGCUCAUUAGGAUAAUUGCAGGCAAUUAACUAAAUUGCCGCCGCAUUUGUCGCU---CCUCG ----((((..(((-((((((((.....((-(........))).....((((..((((((....)))))).)))).))))))))))...)..))))---..... ( -34.30, z-score = -1.39, R) >droYak2.chrX 17226144 96 + 21770863 CAGCGACAUCGG---CUAAAGGCAUUGGC-UAAAGGCAAGGCGCUCAUUAGGAUAAUUGCAGGCAAUUAACUAAAUUGCCGCCGCAUUUGUCGCU---CCUCG .(((((((.(((---(.....((((((.(-(((.(((.....)))..))))..))).))).(((((((.....)))))))))))....)))))))---..... ( -32.60, z-score = -2.03, R) >droEre2.scaffold_4690 1256893 96 - 18748788 CAGCGACAUCGG---CUAAAGGCAUUGGC-UAAAGGCAAGGCGCUCAUUAGGAUAAUUGCAGGCAAUUAACUAAAUUGCCGCCGCAUUUGUCGCU---CCUCG .(((((((.(((---(.....((((((.(-(((.(((.....)))..))))..))).))).(((((((.....)))))))))))....)))))))---..... ( -32.60, z-score = -2.03, R) >droAna3.scaffold_13117 11952 94 + 5790199 ----GACGAUAGCAAGGCAGAGGCAGGAG-CUAAGGCAAGGCGCUCAUUAGGAUAAUUGCAGGCAAUUAACUAAAUUGCCGCCGCAUUUGUCGCG---ACUU- ----(((((..((...((...(((....)-))...))..(((((..............)).(((((((.....))))))))))))..)))))...---....- ( -27.84, z-score = -0.88, R) >droWil1.scaffold_180588 1227136 95 - 1294757 --GCGACAUUGGGAUGGCAAAGCAAAGGA-UAUAGGCAAGGCGCUCAUUAAGAUAAUUGCAGGCAAUUAACUAAAUUGCCGCCGCAUUUGUCGCU---CAA-- --((((((....(.((((...((((....-.((..((.....))..))........)))).(((((((.....))))))))))))...)))))).---...-- ( -29.22, z-score = -1.88, R) >dp4.chrXL_group1e 709962 96 + 12523060 ---GAGUGUUGGCAGGCAGCGACAUCGGC-UAGAGGCAAGGCGCUCAUUAGGAUAAUUGCCUGCAAUUAACUAAAUUGCCGCCGCAUUUGUCGCU---CCUCG ---........(.(((.(((((((.((((-...((((((....((.....))....))))))((((((.....)))))).))))....)))))))---)))). ( -34.00, z-score = -1.40, R) >droPer1.super_14 1802183 96 - 2168203 ---GAGUGUUGGCAGGCAGCGACAUCGGC-UAGAGGCAAGGCGCUCAUUAGGAUAAUUGCCUGCAAUUAACUAAAUUGCCGCCGCAUUUGUCGCU---CCUCG ---........(.(((.(((((((.((((-...((((((....((.....))....))))))((((((.....)))))).))))....)))))))---)))). ( -34.00, z-score = -1.40, R) >droGri2.scaffold_14853 2752815 94 + 10151454 ---------GAGUUAGCUGCGACAUUGGCGCUCAGGCAAGGCGCUCAUUAGGAUAAUUGCAGGCAAUUAACUAAAUUGCCGCCGCAUUUGUCGCUUAGCCUCA ---------(((...(((((((((...((((....))..(((((..............)).(((((((.....))))))))))))...))))))..)))))). ( -34.24, z-score = -1.97, R) >consensus ____GACGUUGGC_GGCAGCGACAUCGGC_UAAAGGCAAGGCGCUCAUUAGGAUAAUUGCAGGCAAUUAACUAAAUUGCCGCCGCAUUUGUCGCU___CCUCG .................................(((...(((.((....))(((((.(((.(((((((.....)))))))...))).))))))))...))).. (-17.01 = -17.79 + 0.78)


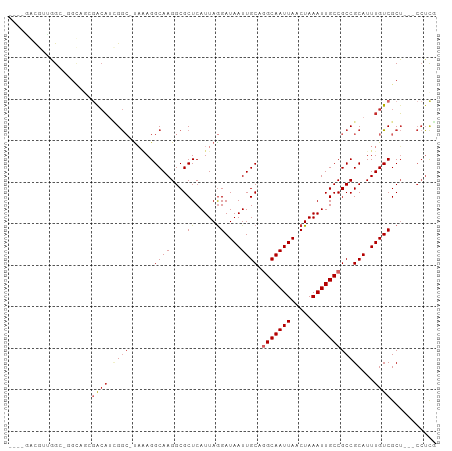
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:13:04 2011